The HIM database manually collected almost all the available in vivo metabolism information for herbal active ingredients, as well as their corresponding bioactivity, organs and/or tissues distribution, toxicity, ADME and the clinical research profile when possible. The paper is: Hong Kang, Kailin Tang, Qi Liu, Yi Sun, Qi Huang, Ruixin Zhu, Jun Gao, Duanfeng Zhang, Chenggang Huang and Zhiwei Cao: HIM-herbal ingredients in-vivo metabolism database. Journal of Cheminformatics 2013, 5(1):28. We are grateful to the authors for creating and curating this resource, and for allowing us to incorporate its structures in ZINC.
We assess the chemical diversity of a subset by clustering the molecules. First, we sort ligands by increasing molecular weight. Then, we use the SUBSET 1.0 algorithm ( Voigt JH, Bienfait B, Wang S, Nicklaus MC. JCICS, 2001, 41, 702-12) to progressively select compounds that differ from those previously selected by at least the Tanimoto cutoff, using ChemAxon default fingerprints. The resulting representatives have two interesting properties:
| Tanimoto Cutoff Level | 60% | 70% | 80% | 90% | 100% |
|---|---|---|---|---|---|
| Number of Representatives | 80 | 110 | 133 | 184 | 663 |
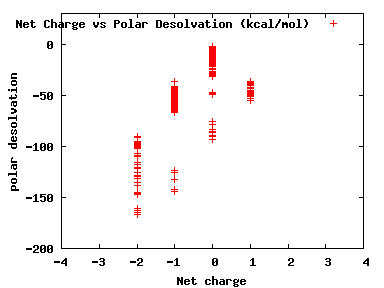
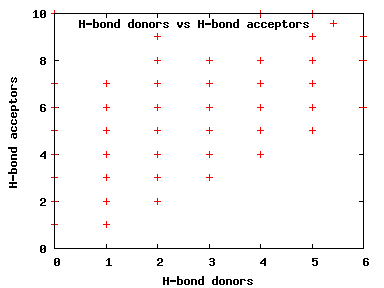
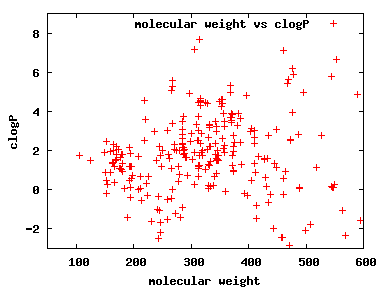
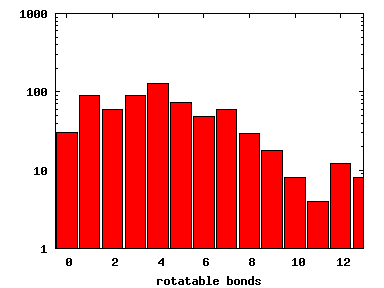
We compute the physical properties of each molecule in the subset, and graph them below.
Download Calculated Physical Properties


| Format | Reference(pH 7) | Mid(pH 6-8) | High(pH 8-9.5) | Low(pH 4.5-6) | Download Unix |
Download Windows |
|---|---|---|---|---|---|---|
| SMILES | All | All | All | All | ||
| MOL2 | All | All | All | All | Single Usual Metals All | Single Usual Metals All |
| SDF | All | All | All | All | Single Usual Metals All | Single Usual Metals All |
| Flexibase | Not Available | Not Available | Not Available | Not Available |