Microbial biocatalytic reactions and biodegradation pathways. The paper is: Ellis LB, Wackett LP. (2012) "Use of the University of Minnesota Biocatalysis/Biodegradation Database for study of microbial degradation" Microbial Informatics and Experimentation 2: 1 (4 January 2012). We are grateful to the authors for creating and curating this database and thank them for allowing us to incorporate its structures in ZINC.
We assess the chemical diversity of a subset by clustering the molecules. First, we sort ligands by increasing molecular weight. Then, we use the SUBSET 1.0 algorithm ( Voigt JH, Bienfait B, Wang S, Nicklaus MC. JCICS, 2001, 41, 702-12) to progressively select compounds that differ from those previously selected by at least the Tanimoto cutoff, using ChemAxon default fingerprints. The resulting representatives have two interesting properties:
| Tanimoto Cutoff Level | 60% | 70% | 80% | 90% | 100% |
|---|---|---|---|---|---|
| Number of Representatives | 158 | 259 | 478 | 714 | 1,517 |
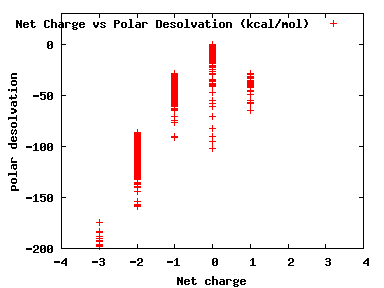
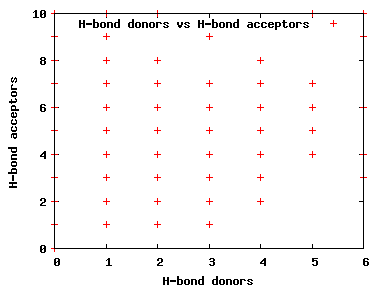
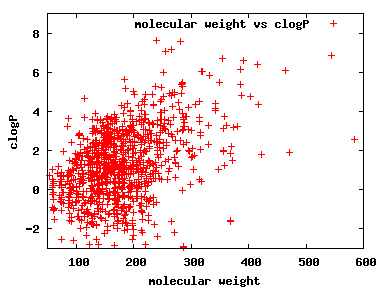
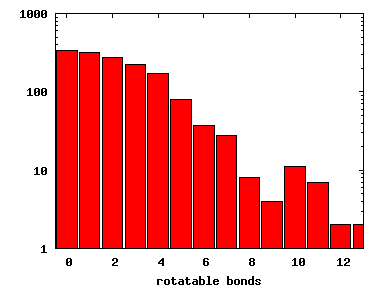
We compute the physical properties of each molecule in the subset, and graph them below.
Download Calculated Physical Properties


| Format | Reference(pH 7) | Mid(pH 6-8) | High(pH 8-9.5) | Low(pH 4.5-6) | Download Unix |
Download Windows |
|---|---|---|---|---|---|---|
| SMILES | All | All | All | All | ||
| MOL2 | All | All | All | All | Single Usual Metals All | Single Usual Metals All |
| SDF | All | All | All | All | Single Usual Metals All | Single Usual Metals All |
| Flexibase | All | All | All | All | Single Usual Metals All | Single Usual Metals All |