|
|
|
|
|
Analogs
-
4760617
-
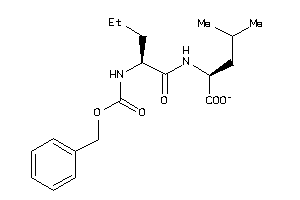
-
4598914
-
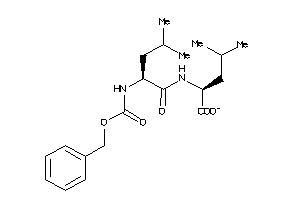
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7 |
0.42 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.52 |
-1.01 |
-51.39 |
2 |
7 |
-1 |
107 |
377.461 |
11 |
↓
|
|
|
|
Analogs
-
7998874
-
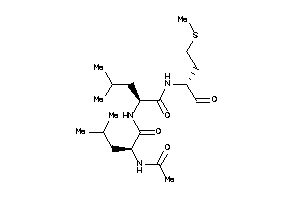
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
23 |
0.40 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.55 |
6.93 |
-21.16 |
3 |
7 |
0 |
104 |
401.573 |
13 |
↓
|
|
|
|
|
|
|
Analogs
-
1544687
-
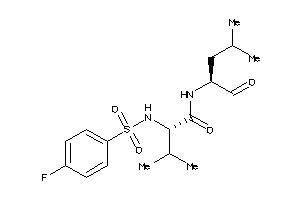
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
8 |
0.45 |
Binding ≤ 10μM
|
|
CATB-1-E |
Cathepsin B (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
78 |
0.40 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.85 |
4.47 |
-14.12 |
2 |
6 |
0 |
92 |
372.462 |
9 |
↓
|
|
|
|
|
|
|
Analogs
-
38255727
-
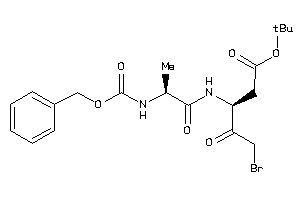
Draw
Identity
99%
90%
80%
70%
Vendors
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.84 |
6.72 |
-17.47 |
2 |
8 |
0 |
111 |
396.415 |
12 |
↓
|
|
|
|
Analogs
-
38255727
-

Draw
Identity
99%
90%
80%
70%
Vendors
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.84 |
6.78 |
-19.28 |
2 |
8 |
0 |
111 |
396.415 |
12 |
↓
|
|
|
|
|
|
|
|
|
|
Analogs
-
606754
-
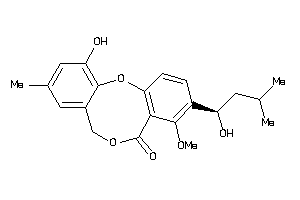
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7100 |
0.27 |
Binding ≤ 10μM
|
|
CAN2-1-E |
Calpain 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
7100 |
0.27 |
Binding ≤ 10μM
|
|
CPNS1-1-E |
Calpain Small Subunit 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7100 |
0.27 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.97 |
5.64 |
-13.07 |
2 |
6 |
0 |
85 |
372.417 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
3.97 |
6.51 |
-53.76 |
1 |
6 |
-1 |
88 |
371.409 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
600 |
0.31 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.79 |
6.27 |
-29.95 |
3 |
6 |
0 |
98 |
391.814 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.79 |
7.06 |
-48.96 |
2 |
6 |
-1 |
101 |
390.806 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.79 |
6.71 |
-67.67 |
4 |
6 |
1 |
99 |
392.822 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-2-E |
Calpain 1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
400 |
0.69 |
Binding ≤ 10μM
|
|
CAN2-1-E |
Calpain 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
400 |
0.69 |
Binding ≤ 10μM
|
|
CPNS1-2-E |
Calpain Small Subunit 1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
400 |
0.69 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.88 |
-1.78 |
-42.84 |
0 |
2 |
-1 |
40 |
305.116 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1300 |
0.48 |
Binding ≤ 10μM
|
|
CAN2-1-E |
Calpain 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1300 |
0.48 |
Binding ≤ 10μM
|
|
CPNS1-1-E |
Calpain Small Subunit 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1300 |
0.48 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.16 |
-0.07 |
-9.41 |
1 |
3 |
0 |
45 |
228.295 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
41 |
0.40 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.52 |
8.15 |
-11.63 |
2 |
6 |
0 |
85 |
362.47 |
11 |
↓
|
|
|
|
Analogs
-
1996112
-
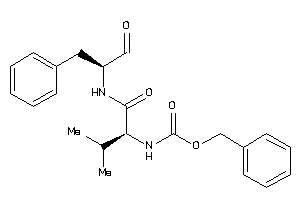
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
20 |
0.38 |
Binding ≤ 10μM
|
|
CAN2-1-E |
Calpain 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
10 |
0.40 |
Binding ≤ 10μM
|
|
CANX-1-E |
Calpain 1/2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
10 |
0.40 |
Binding ≤ 10μM
|
|
CATB-1-E |
Cathepsin B (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
25 |
0.38 |
Binding ≤ 10μM
|
|
CATL1-1-E |
Cathepsin L (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
1270 |
0.29 |
Binding ≤ 10μM
|
|
CPNS1-1-E |
Calpain Small Subunit 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
40 |
0.37 |
Binding ≤ 10μM |
|
Q3HTL5-1-E |
Falcipain 2B (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
5000 |
0.27 |
Binding ≤ 10μM
|
|
Q9N6S8-1-E |
Falcipain 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
5000 |
0.27 |
Binding ≤ 10μM
|
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.31 |
Functional ≤ 10μM
|
|
CAN2-1-E |
Calpain 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.31 |
Functional ≤ 10μM
|
|
CPNS1-1-E |
Calpain Small Subunit 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.31 |
Functional ≤ 10μM
|
|
Z100494-1-O |
C2C12 (Myoblast Cells) (cluster #1 Of 1), Other |
Other |
10000 |
0.25 |
Binding ≤ 10μM
|
|
Z50425-3-O |
Plasmodium Falciparum (cluster #3 Of 22), Other |
Other |
3981 |
0.27 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.17 |
8.75 |
-13.99 |
2 |
6 |
0 |
85 |
382.46 |
10 |
↓
|
|
|
|
Analogs
-
1534489
-
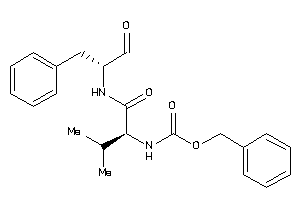
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
20 |
0.38 |
Binding ≤ 10μM
|
|
CAN2-1-E |
Calpain 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
10 |
0.40 |
Binding ≤ 10μM
|
|
CANX-1-E |
Calpain 1/2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
10 |
0.40 |
Binding ≤ 10μM
|
|
CATB-1-E |
Cathepsin B (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
25 |
0.38 |
Binding ≤ 10μM
|
|
CATL1-1-E |
Cathepsin L (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
1270 |
0.29 |
Binding ≤ 10μM
|
|
CPNS1-1-E |
Calpain Small Subunit 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
40 |
0.37 |
Binding ≤ 10μM |
|
Q3HTL5-1-E |
Falcipain 2B (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
5000 |
0.27 |
Binding ≤ 10μM
|
|
Q9N6S8-1-E |
Falcipain 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
5000 |
0.27 |
Binding ≤ 10μM
|
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.31 |
Functional ≤ 10μM
|
|
CAN2-1-E |
Calpain 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.31 |
Functional ≤ 10μM
|
|
CPNS1-1-E |
Calpain Small Subunit 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.31 |
Functional ≤ 10μM
|
|
Z100494-1-O |
C2C12 (Myoblast Cells) (cluster #1 Of 1), Other |
Other |
10000 |
0.25 |
Binding ≤ 10μM
|
|
Z50425-3-O |
Plasmodium Falciparum (cluster #3 Of 22), Other |
Other |
3981 |
0.27 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.17 |
8.75 |
-13.21 |
2 |
6 |
0 |
85 |
382.46 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 15 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
10 |
0.59 |
Binding ≤ 10μM
|
|
CPNS1-1-E |
Calpain Small Subunit 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
10 |
0.59 |
Binding ≤ 10μM
|
|
S15A2-1-E |
Solute Carrier Family 15 Member 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
3800 |
0.40 |
ADME/T ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.01 |
-2.02 |
-70.32 |
4 |
5 |
0 |
96 |
264.325 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 5 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CAN1-1-E |
Calpain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7 |
0.44 |
Binding ≤ 10μM
|
|
CATK-1-E |
Cathepsin K (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.84 |
8.3 |
-11.2 |
2 |
6 |
0 |
85 |
362.47 |
12 |
↓
|
|