|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.93 |
12.79 |
-11.95 |
2 |
8 |
0 |
86 |
432.528 |
9 |
↓
|
|
Lo
Low (pH 4.5-6)
|
4.93 |
13.07 |
-35.61 |
3 |
8 |
1 |
87 |
433.536 |
9 |
↓
|
|
Lo
Low (pH 4.5-6)
|
4.93 |
13.18 |
-31.89 |
3 |
8 |
1 |
87 |
433.536 |
9 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 75 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYSY-3-E |
Thymidylate Synthase (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
7000 |
0.80 |
Binding ≤ 10μM
|
|
Z103205-4-O |
A431 (cluster #4 Of 4), Other |
Other |
18 |
1.20 |
Functional ≤ 10μM
|
|
Z103419-1-O |
UACC-812 (cluster #1 Of 2), Other |
Other |
3100 |
0.86 |
Functional ≤ 10μM
|
|
Z50425-14-O |
Plasmodium Falciparum (cluster #14 Of 22), Other |
Other |
7943 |
0.79 |
Functional ≤ 10μM
|
|
Z80035-6-O |
B16 (Melanoma Cells) (cluster #6 Of 7), Other |
Other |
8730 |
0.79 |
Functional ≤ 10μM
|
|
Z80054-2-O |
Caco-2 (Colon Adenocarcinoma Cells) (cluster #2 Of 4), Other |
Other |
1500 |
0.91 |
Functional ≤ 10μM
|
|
Z80064-6-O |
CCRF-CEM (T-cell Leukemia) (cluster #6 Of 9), Other |
Other |
5000 |
0.82 |
Functional ≤ 10μM
|
|
Z80088-4-O |
CHO (Ovarian Cells) (cluster #4 Of 4), Other |
Other |
5000 |
0.82 |
Functional ≤ 10μM
|
|
Z80101-1-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #1 Of 2), Other |
Other |
1100 |
0.93 |
Functional ≤ 10μM |
|
Z80120-3-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
6000 |
0.81 |
Functional ≤ 10μM |
|
Z80125-6-O |
DU-145 (Prostate Carcinoma) (cluster #6 Of 9), Other |
Other |
4800 |
0.83 |
Functional ≤ 10μM
|
|
Z80136-3-O |
FM3A (Breast Carcinoma Cells) (cluster #3 Of 6), Other |
Other |
180 |
1.05 |
Functional ≤ 10μM
|
|
Z80156-6-O |
HL-60 (Promyeloblast Leukemia Cells) (cluster #6 Of 12), Other |
Other |
9500 |
0.78 |
Functional ≤ 10μM
|
|
Z80164-4-O |
HT-1080 (Fibrosarcoma Cells) (cluster #4 Of 6), Other |
Other |
9450 |
0.78 |
Functional ≤ 10μM
|
|
Z80166-8-O |
HT-29 (Colon Adenocarcinoma Cells) (cluster #8 Of 12), Other |
Other |
7300 |
0.80 |
Functional ≤ 10μM
|
|
Z80186-7-O |
K562 (Erythroleukemia Cells) (cluster #7 Of 11), Other |
Other |
2400 |
0.87 |
Functional ≤ 10μM
|
|
Z80192-2-O |
KM12 (Colon Adenocarcinoma Cells) (cluster #2 Of 4), Other |
Other |
8320 |
0.79 |
Functional ≤ 10μM
|
|
Z80193-8-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #8 Of 12), Other |
Other |
970 |
0.94 |
Functional ≤ 10μM
|
|
Z80224-5-O |
MCF7 (Breast Carcinoma Cells) (cluster #5 Of 14), Other |
Other |
9600 |
0.78 |
Functional ≤ 10μM
|
|
Z80268-1-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
2500 |
0.87 |
Functional ≤ 10μM |
|
Z80269-3-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #3 Of 3), Other |
Other |
3000 |
0.86 |
Functional ≤ 10μM |
|
Z80347-2-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
4400 |
0.83 |
Functional ≤ 10μM |
|
Z80362-5-O |
P388 (Lymphoma Cells) (cluster #5 Of 8), Other |
Other |
609 |
0.97 |
Functional ≤ 10μM |
|
Z80390-6-O |
PC-3 (Prostate Carcinoma Cells) (cluster #6 Of 10), Other |
Other |
16 |
1.21 |
Functional ≤ 10μM
|
|
Z80485-4-O |
SK-MEL-28 (Melanoma Cells) (cluster #4 Of 6), Other |
Other |
4500 |
0.83 |
Functional ≤ 10μM
|
|
Z80490-4-O |
SK-MES-1 (cluster #4 Of 4), Other |
Other |
310 |
1.01 |
Functional ≤ 10μM
|
|
Z80525-2-O |
SW48 (cluster #2 Of 2), Other |
Other |
5700 |
0.82 |
Functional ≤ 10μM |
|
Z80526-3-O |
SW480 (Colon Adenocarcinoma Cells) (cluster #3 Of 6), Other |
Other |
8100 |
0.79 |
Functional ≤ 10μM
|
|
Z80532-5-O |
T-24 (Bladder Carcinoma Cells) (cluster #5 Of 6), Other |
Other |
4100 |
0.84 |
Functional ≤ 10μM
|
|
Z80583-6-O |
Vero (Kidney Cells) (cluster #6 Of 7), Other |
Other |
6810 |
0.80 |
Functional ≤ 10μM
|
|
Z80641-1-O |
791T Cell Line (cluster #1 Of 1), Other |
Other |
5500 |
0.82 |
Functional ≤ 10μM
|
|
Z80653-2-O |
9L (Glioma Cells) (cluster #2 Of 2), Other |
Other |
1000 |
0.93 |
Functional ≤ 10μM
|
|
Z80682-9-O |
A549 (Lung Carcinoma Cells) (cluster #9 Of 11), Other |
Other |
9200 |
0.78 |
Functional ≤ 10μM
|
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
3900 |
0.84 |
Functional ≤ 10μM |
|
Z80712-5-O |
T47D (Breast Carcinoma Cells) (cluster #5 Of 7), Other |
Other |
600 |
0.97 |
Functional ≤ 10μM
|
|
Z80724-1-O |
C170 Cell Line (cluster #1 Of 1), Other |
Other |
2000 |
0.89 |
Functional ≤ 10μM
|
|
Z80742-2-O |
C6 (Glioma Cells) (cluster #2 Of 3), Other |
Other |
6100 |
0.81 |
Functional ≤ 10μM
|
|
Z80874-5-O |
CEM (T-cell Leukemia) (cluster #5 Of 7), Other |
Other |
9230 |
0.78 |
Functional ≤ 10μM
|
|
Z80928-7-O |
HCT-116 (Colon Carcinoma Cells) (cluster #7 Of 9), Other |
Other |
8510 |
0.79 |
Functional ≤ 10μM
|
|
Z81020-1-O |
HepG2 (Hepatoblastoma Cells) (cluster #1 Of 8), Other |
Other |
9200 |
0.78 |
Functional ≤ 10μM
|
|
Z81024-6-O |
NCI-H460 (Non-small Cell Lung Carcinoma) (cluster #6 Of 8), Other |
Other |
8500 |
0.79 |
Functional ≤ 10μM |
|
Z81034-5-O |
A2780 (Ovarian Carcinoma Cells) (cluster #5 Of 10), Other |
Other |
7210 |
0.80 |
Functional ≤ 10μM
|
|
Z81041-1-O |
Prostatic Carcinoma Cells (cluster #1 Of 2), Other |
Other |
6400 |
0.81 |
Functional ≤ 10μM
|
|
Z81043-2-O |
N592 (cluster #2 Of 2), Other |
Other |
5400 |
0.82 |
Functional ≤ 10μM
|
|
Z81048-1-O |
TSU (Prostatic Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
3600 |
0.85 |
Functional ≤ 10μM
|
|
Z81115-6-O |
KB (Squamous Cell Carcinoma) (cluster #6 Of 6), Other |
Other |
4460 |
0.83 |
Functional ≤ 10μM
|
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
6200 |
0.81 |
Functional ≤ 10μM |
|
Z81170-4-O |
LNCaP (Prostate Carcinoma) (cluster #4 Of 5), Other |
Other |
4900 |
0.83 |
Functional ≤ 10μM
|
|
Z81247-7-O |
HeLa (Cervical Adenocarcinoma Cells) (cluster #7 Of 9), Other |
Other |
9700 |
0.78 |
Functional ≤ 10μM
|
|
Z81252-9-O |
MDA-MB-231 (Breast Adenocarcinoma Cells) (cluster #9 Of 11), Other |
Other |
9600 |
0.78 |
Functional ≤ 10μM
|
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
2700 |
0.87 |
Functional ≤ 10μM |
|
Z81331-3-O |
SW-620 (Colon Adenocarcinoma Cells) (cluster #3 Of 6), Other |
Other |
9000 |
0.78 |
Functional ≤ 10μM
|
|
Z81335-3-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
9710 |
0.78 |
Functional ≤ 10μM
|
|
Z80156-3-O |
HL-60 (Promyeloblast Leukemia Cells) (cluster #3 Of 4), Other |
Other |
160 |
1.06 |
ADME/T ≤ 10μM
|
|
Z80192-2-O |
KM12 (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
7940 |
0.79 |
ADME/T ≤ 10μM
|
|
Z80193-4-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #4 Of 4), Other |
Other |
560 |
0.97 |
ADME/T ≤ 10μM
|
|
Z80583-3-O |
Vero (Kidney Cells) (cluster #3 Of 3), Other |
Other |
200 |
1.04 |
ADME/T ≤ 10μM
|
|
Z80901-2-O |
HaCaT (Keratinocytes) (cluster #2 Of 2), Other |
Other |
400 |
1.00 |
ADME/T ≤ 10μM
|
|
Z80928-2-O |
HCT-116 (Colon Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
2040 |
0.89 |
ADME/T ≤ 10μM
|
|
Z81247-2-O |
HeLa (Cervical Adenocarcinoma Cells) (cluster #2 Of 3), Other |
Other |
570 |
0.97 |
ADME/T ≤ 10μM
|
|
Z81335-2-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
6610 |
0.81 |
ADME/T ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.59 |
-1.23 |
-13.33 |
2 |
4 |
0 |
66 |
130.078 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
-0.13 |
-4 |
-48.58 |
1 |
4 |
-1 |
69 |
129.07 |
0 |
↓
|
|
Mid
Mid (pH 6-8)
|
-0.13 |
-3.14 |
-32.57 |
1 |
4 |
-1 |
69 |
129.07 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CDC7-1-E |
Cell Division Cycle 7-related Protein Kinase (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
10 |
0.70 |
Binding ≤ 10μM
|
|
CDK1-1-E |
Cyclin-dependent Kinase 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
250 |
0.58 |
Binding ≤ 10μM
|
|
CDK2-1-E |
Cyclin-dependent Kinase 2 (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
240 |
0.58 |
Binding ≤ 10μM
|
|
CDK5-1-E |
Cyclin-dependent Kinase 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
460 |
0.55 |
Binding ≤ 10μM
|
|
CDK9-1-E |
Cyclin-dependent Kinase 9 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
34 |
0.65 |
Binding ≤ 10μM
|
|
CHK2-1-E |
Serine/threonine-protein Kinase Chk2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1100 |
0.52 |
Binding ≤ 10μM
|
|
GSK3B-1-E |
Glycogen Synthase Kinase-3 Beta (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
220 |
0.58 |
Binding ≤ 10μM
|
|
KS6A4-1-E |
Ribosomal Protein S6 Kinase Alpha 4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
3010 |
0.48 |
Binding ≤ 10μM
|
|
KS6A5-2-E |
Ribosomal Protein S6 Kinase Alpha 5 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
2340 |
0.49 |
Binding ≤ 10μM
|
|
MAPK2-1-E |
MAP Kinase-activated Protein Kinase 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
470 |
0.55 |
Binding ≤ 10μM
|
|
MAPK5-1-E |
MAP Kinase-activated Protein Kinase 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
500 |
0.55 |
Binding ≤ 10μM
|
|
MKNK1-1-E |
MAP Kinase-interacting Serine/threonine-protein Kinase MNK1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
2670 |
0.49 |
Binding ≤ 10μM
|
|
MKNK2-1-E |
MAP Kinase Signal-integrating Kinase 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
171 |
0.59 |
Binding ≤ 10μM
|
|
PLK1-1-E |
Serine/threonine-protein Kinase PLK1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
980 |
0.53 |
Binding ≤ 10μM
|
|
Z80166-1-O |
HT-29 (Colon Adenocarcinoma Cells) (cluster #1 Of 12), Other |
Other |
5000 |
0.46 |
Functional ≤ 10μM
|
|
Z80186-1-O |
K562 (Erythroleukemia Cells) (cluster #1 Of 11), Other |
Other |
5870 |
0.46 |
Functional ≤ 10μM
|
|
Z80224-1-O |
MCF7 (Breast Carcinoma Cells) (cluster #1 Of 14), Other |
Other |
1330 |
0.51 |
Functional ≤ 10μM
|
|
Z80390-1-O |
PC-3 (Prostate Carcinoma Cells) (cluster #1 Of 10), Other |
Other |
4590 |
0.47 |
Functional ≤ 10μM
|
|
Z80466-1-O |
SF268 (cluster #1 Of 4), Other |
Other |
860 |
0.53 |
Functional ≤ 10μM
|
|
Z80525-1-O |
SW48 (cluster #1 Of 2), Other |
Other |
1200 |
0.52 |
Functional ≤ 10μM
|
|
Z80526-1-O |
SW480 (Colon Adenocarcinoma Cells) (cluster #1 Of 6), Other |
Other |
2670 |
0.49 |
Functional ≤ 10μM
|
|
Z80561-1-O |
U2OS (Osteosarcoma Cells) (cluster #1 Of 1), Other |
Other |
1490 |
0.51 |
Functional ≤ 10μM
|
|
Z80928-3-O |
HCT-116 (Colon Carcinoma Cells) (cluster #3 Of 9), Other |
Other |
970 |
0.53 |
Functional ≤ 10μM
|
|
Z81034-3-O |
A2780 (Ovarian Carcinoma Cells) (cluster #3 Of 10), Other |
Other |
1100 |
0.52 |
Functional ≤ 10μM
|
|
Z81072-1-O |
Jurkat (Acute Leukemic T-cells) (cluster #1 Of 10), Other |
Other |
3200 |
0.48 |
Functional ≤ 10μM
|
|
Z81247-1-O |
HeLa (Cervical Adenocarcinoma Cells) (cluster #1 Of 9), Other |
Other |
1830 |
0.50 |
Functional ≤ 10μM
|
|
Z81335-1-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #1 Of 5), Other |
Other |
3810 |
0.47 |
Functional ≤ 10μM
|
|
Z80335-1-O |
NHDF (cluster #1 Of 2), Other |
Other |
1600 |
0.51 |
ADME/T ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.64 |
2.58 |
-15.17 |
2 |
4 |
0 |
58 |
213.24 |
1 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.64 |
2.88 |
-44.62 |
3 |
4 |
1 |
59 |
214.248 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 19 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
HDA10-1-E |
Histone Deacetylase 10 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.27 |
Binding ≤ 10μM
|
|
HDA11-1-E |
Histone Deacetylase 11 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.27 |
Binding ≤ 10μM
|
|
HDAC1-1-E |
Histone Deacetylase 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
600 |
0.31 |
Binding ≤ 10μM
|
|
HDAC2-2-E |
Histone Deacetylase 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
65 |
0.36 |
Binding ≤ 10μM
|
|
HDAC3-1-E |
Histone Deacetylase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
740 |
0.31 |
Binding ≤ 10μM
|
|
HDAC4-1-E |
Histone Deacetylase 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
510 |
0.31 |
Binding ≤ 10μM
|
|
HDAC5-1-E |
Histone Deacetylase 5 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.27 |
Binding ≤ 10μM
|
|
HDAC6-1-E |
Histone Deacetylase 6 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.27 |
Binding ≤ 10μM
|
|
HDAC7-1-E |
Histone Deacetylase 7 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.27 |
Binding ≤ 10μM
|
|
HDAC8-1-E |
Histone Deacetylase 8 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.27 |
Binding ≤ 10μM
|
|
HDAC9-1-E |
Histone Deacetylase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.27 |
Binding ≤ 10μM
|
|
Q9XYC7-1-E |
Histone Deacetylase (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
940 |
0.30 |
Binding ≤ 10μM
|
|
TF-1-E |
Coagulation Factor III (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
6000 |
0.26 |
Binding ≤ 10μM
|
|
HDA10-1-E |
Histone Deacetylase 10 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDA11-1-E |
Histone Deacetylase 11 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC1-1-E |
Histone Deacetylase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC2-1-E |
Histone Deacetylase 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC3-1-E |
Histone Deacetylase 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC4-1-E |
Histone Deacetylase 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC5-1-E |
Histone Deacetylase 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC6-1-E |
Histone Deacetylase 6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC7-1-E |
Histone Deacetylase 7 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC8-1-E |
Histone Deacetylase 8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
HDAC9-1-E |
Histone Deacetylase 9 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.30 |
Functional ≤ 10μM
|
|
CP3A4-2-E |
Cytochrome P450 3A4 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
730 |
0.31 |
ADME/T ≤ 10μM |
|
Z80125-1-O |
DU-145 (Prostate Carcinoma) (cluster #1 Of 9), Other |
Other |
2000 |
0.28 |
Functional ≤ 10μM
|
|
Z80224-1-O |
MCF7 (Breast Carcinoma Cells) (cluster #1 Of 14), Other |
Other |
5000 |
0.27 |
Functional ≤ 10μM
|
|
Z80269-1-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #1 Of 3), Other |
Other |
4160 |
0.27 |
Functional ≤ 10μM
|
|
Z80475-2-O |
SK-BR-3 (Breast Adenocarcinoma) (cluster #2 Of 3), Other |
Other |
870 |
0.30 |
Functional ≤ 10μM
|
|
Z80525-1-O |
SW48 (cluster #1 Of 2), Other |
Other |
5000 |
0.27 |
Functional ≤ 10μM
|
|
Z80532-3-O |
T-24 (Bladder Carcinoma Cells) (cluster #3 Of 6), Other |
Other |
2500 |
0.28 |
Functional ≤ 10μM
|
|
Z80682-2-O |
A549 (Lung Carcinoma Cells) (cluster #2 Of 11), Other |
Other |
8000 |
0.25 |
Functional ≤ 10μM
|
|
Z80928-3-O |
HCT-116 (Colon Carcinoma Cells) (cluster #3 Of 9), Other |
Other |
800 |
0.30 |
Functional ≤ 10μM
|
|
Z81020-4-O |
HepG2 (Hepatoblastoma Cells) (cluster #4 Of 8), Other |
Other |
1 |
0.45 |
Functional ≤ 10μM
|
|
Z81034-3-O |
A2780 (Ovarian Carcinoma Cells) (cluster #3 Of 10), Other |
Other |
3000 |
0.28 |
Functional ≤ 10μM
|
|
Z81170-1-O |
LNCaP (Prostate Carcinoma) (cluster #1 Of 5), Other |
Other |
360 |
0.32 |
Functional ≤ 10μM
|
|
Z81199-1-O |
NCI-H446 (Small Cell Lung Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
1000 |
0.30 |
Functional ≤ 10μM
|
|
Z81247-1-O |
HeLa (Cervical Adenocarcinoma Cells) (cluster #1 Of 9), Other |
Other |
1800 |
0.29 |
Functional ≤ 10μM
|
|
Z81252-7-O |
MDA-MB-231 (Breast Adenocarcinoma Cells) (cluster #7 Of 11), Other |
Other |
3000 |
0.28 |
Functional ≤ 10μM
|
|
Z81331-1-O |
SW-620 (Colon Adenocarcinoma Cells) (cluster #1 Of 6), Other |
Other |
3200 |
0.27 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.03 |
5.3 |
-15.7 |
4 |
7 |
0 |
106 |
376.416 |
7 |
↓
|
|
Lo
Low (pH 4.5-6)
|
2.03 |
5.76 |
-45.32 |
5 |
7 |
1 |
108 |
377.424 |
7 |
↓
|
|
|
|
Analogs
-
5828039
-
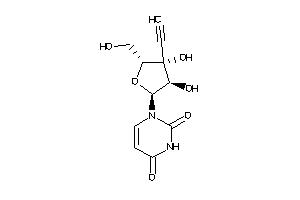
-
5828050
-
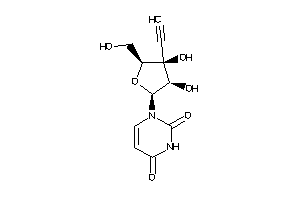
-
5828069
-
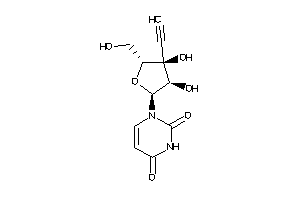
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z80101-2-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
60 |
0.53 |
Functional ≤ 10μM |
|
Z80120-2-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #2 Of 5), Other |
Other |
66 |
0.53 |
Functional ≤ 10μM |
|
Z80164-1-O |
HT-1080 (Fibrosarcoma Cells) (cluster #1 Of 6), Other |
Other |
24 |
0.56 |
Functional ≤ 10μM |
|
Z80268-2-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
80 |
0.52 |
Functional ≤ 10μM |
|
Z80269-1-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #1 Of 3), Other |
Other |
24 |
0.56 |
Functional ≤ 10μM |
|
Z80347-1-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
34 |
0.55 |
Functional ≤ 10μM |
|
Z80525-1-O |
SW48 (cluster #1 Of 2), Other |
Other |
120 |
0.51 |
Functional ≤ 10μM |
|
Z80532-1-O |
T-24 (Bladder Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
34 |
0.55 |
Functional ≤ 10μM |
|
Z80682-1-O |
A549 (Lung Carcinoma Cells) (cluster #1 Of 11), Other |
Other |
29 |
0.56 |
Functional ≤ 10μM |
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
36 |
0.55 |
Functional ≤ 10μM |
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
190 |
0.50 |
Functional ≤ 10μM |
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
18 |
0.57 |
Functional ≤ 10μM |
|
Z81335-2-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #2 Of 5), Other |
Other |
43 |
0.54 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.82 |
-8.58 |
-11.81 |
4 |
8 |
0 |
124 |
268.225 |
2 |
↓
|
|
|
|
Analogs
-
5828050
-

-
5828069
-

-
5828032
-
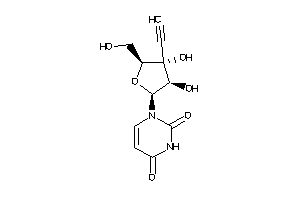
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z80101-2-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
60 |
0.53 |
Functional ≤ 10μM |
|
Z80120-2-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #2 Of 5), Other |
Other |
66 |
0.53 |
Functional ≤ 10μM |
|
Z80164-1-O |
HT-1080 (Fibrosarcoma Cells) (cluster #1 Of 6), Other |
Other |
24 |
0.56 |
Functional ≤ 10μM |
|
Z80268-2-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
80 |
0.52 |
Functional ≤ 10μM |
|
Z80269-1-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #1 Of 3), Other |
Other |
24 |
0.56 |
Functional ≤ 10μM |
|
Z80347-1-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
34 |
0.55 |
Functional ≤ 10μM |
|
Z80525-1-O |
SW48 (cluster #1 Of 2), Other |
Other |
120 |
0.51 |
Functional ≤ 10μM |
|
Z80532-1-O |
T-24 (Bladder Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
34 |
0.55 |
Functional ≤ 10μM |
|
Z80682-1-O |
A549 (Lung Carcinoma Cells) (cluster #1 Of 11), Other |
Other |
29 |
0.56 |
Functional ≤ 10μM |
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
36 |
0.55 |
Functional ≤ 10μM |
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
190 |
0.50 |
Functional ≤ 10μM |
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
18 |
0.57 |
Functional ≤ 10μM |
|
Z81335-2-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #2 Of 5), Other |
Other |
43 |
0.54 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.82 |
-8.19 |
-11.61 |
4 |
8 |
0 |
124 |
268.225 |
2 |
↓
|
|
|
|
Analogs
-
4265274
-
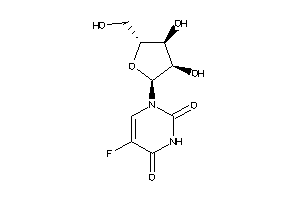
-
6090974
-
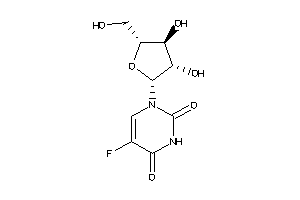
-
12651207
-
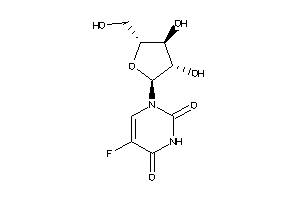
Draw
Identity
99%
90%
80%
70%
Vendors
And 20 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z80064-1-O |
CCRF-CEM (T-cell Leukemia) (cluster #1 Of 9), Other |
Other |
50 |
0.57 |
Functional ≤ 10μM
|
|
Z80101-2-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
5 |
0.65 |
Functional ≤ 10μM |
|
Z80120-1-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #1 Of 5), Other |
Other |
73 |
0.56 |
Functional ≤ 10μM |
|
Z80136-1-O |
FM3A (Breast Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
14 |
0.61 |
Functional ≤ 10μM
|
|
Z80164-1-O |
HT-1080 (Fibrosarcoma Cells) (cluster #1 Of 6), Other |
Other |
41 |
0.57 |
Functional ≤ 10μM |
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
5 |
0.65 |
Functional ≤ 10μM
|
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
720 |
0.48 |
Functional ≤ 10μM
|
|
Z80268-2-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
28 |
0.59 |
Functional ≤ 10μM |
|
Z80269-1-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #1 Of 3), Other |
Other |
26 |
0.59 |
Functional ≤ 10μM |
|
Z80347-2-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
22 |
0.60 |
Functional ≤ 10μM |
|
Z80525-2-O |
SW48 (cluster #2 Of 2), Other |
Other |
16 |
0.61 |
Functional ≤ 10μM |
|
Z80532-1-O |
T-24 (Bladder Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
14 |
0.61 |
Functional ≤ 10μM |
|
Z80682-9-O |
A549 (Lung Carcinoma Cells) (cluster #9 Of 11), Other |
Other |
1900 |
0.44 |
Functional ≤ 10μM |
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
50 |
0.57 |
Functional ≤ 10μM |
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
16 |
0.61 |
Functional ≤ 10μM |
|
Z81247-7-O |
HeLa (Cervical Adenocarcinoma Cells) (cluster #7 Of 9), Other |
Other |
22 |
0.60 |
Functional ≤ 10μM
|
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
45 |
0.57 |
Functional ≤ 10μM |
|
Z81335-3-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
47 |
0.57 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.68 |
-6.46 |
-14.66 |
4 |
8 |
0 |
125 |
262.193 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.22 |
-9.21 |
-53.85 |
3 |
8 |
-1 |
128 |
261.185 |
2 |
↓
|
|
|
|
Analogs
-
388678
-
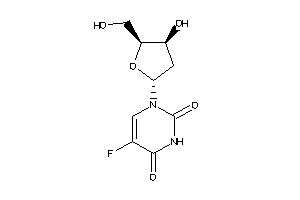
-
489135
-
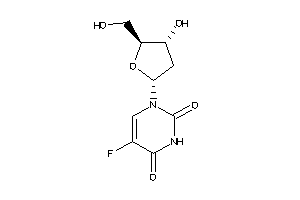
-
3813010
-
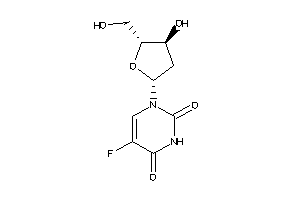
-
3830846
-
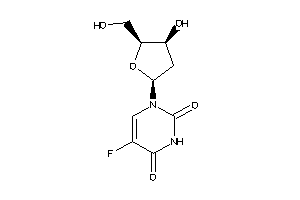
-
3977743
-
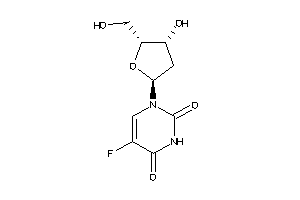
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYSY-1-E |
Thymidylate Synthase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
7300 |
0.42 |
Binding ≤ 10μM
|
|
Z80064-6-O |
CCRF-CEM (T-cell Leukemia) (cluster #6 Of 9), Other |
Other |
500 |
0.52 |
Functional ≤ 10μM
|
|
Z80101-2-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
650 |
0.51 |
Functional ≤ 10μM |
|
Z80120-3-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
92 |
0.58 |
Functional ≤ 10μM |
|
Z80164-4-O |
HT-1080 (Fibrosarcoma Cells) (cluster #4 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
45 |
0.61 |
Functional ≤ 10μM
|
|
Z80208-1-O |
L5178Y (Lymphoma Cells) (cluster #1 Of 2), Other |
Other |
2 |
0.72 |
Functional ≤ 10μM
|
|
Z80268-2-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80269-3-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #3 Of 3), Other |
Other |
3500 |
0.45 |
Functional ≤ 10μM |
|
Z80347-2-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
15 |
0.64 |
Functional ≤ 10μM |
|
Z80525-2-O |
SW48 (cluster #2 Of 2), Other |
Other |
130 |
0.57 |
Functional ≤ 10μM |
|
Z80532-1-O |
T-24 (Bladder Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80682-9-O |
A549 (Lung Carcinoma Cells) (cluster #9 Of 11), Other |
Other |
1000 |
0.49 |
Functional ≤ 10μM |
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
50 |
0.60 |
Functional ≤ 10μM |
|
Z81048-1-O |
TSU (Prostatic Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
58 |
0.60 |
Functional ≤ 10μM
|
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
760 |
0.50 |
Functional ≤ 10μM |
|
Z81137-1-O |
L929 (Fibroblast Cells) (cluster #1 Of 2), Other |
Other |
7700 |
0.42 |
Functional ≤ 10μM
|
|
Z81170-4-O |
LNCaP (Prostate Carcinoma) (cluster #4 Of 5), Other |
Other |
69 |
0.59 |
Functional ≤ 10μM
|
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
3 |
0.70 |
Functional ≤ 10μM |
|
Z81335-3-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
49 |
0.60 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.72 |
-5.12 |
-11.33 |
3 |
7 |
0 |
105 |
246.194 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.26 |
-7.87 |
-48.56 |
2 |
7 |
-1 |
108 |
245.186 |
2 |
↓
|
|
|
|
Analogs
-
3977743
-

-
3977744
-
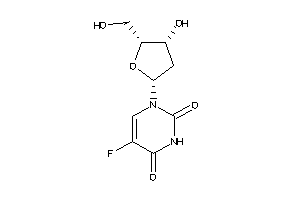
-
4045293
-
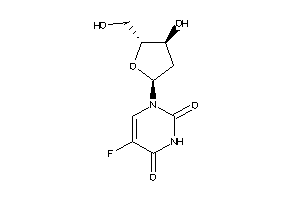
-
1457
-
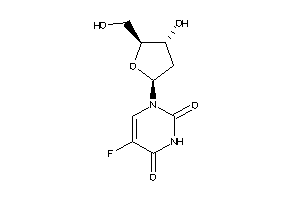
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYSY-1-E |
Thymidylate Synthase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
7300 |
0.42 |
Binding ≤ 10μM
|
|
Z80064-6-O |
CCRF-CEM (T-cell Leukemia) (cluster #6 Of 9), Other |
Other |
500 |
0.52 |
Functional ≤ 10μM
|
|
Z80101-2-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
650 |
0.51 |
Functional ≤ 10μM |
|
Z80120-3-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
92 |
0.58 |
Functional ≤ 10μM |
|
Z80164-4-O |
HT-1080 (Fibrosarcoma Cells) (cluster #4 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
45 |
0.61 |
Functional ≤ 10μM
|
|
Z80208-1-O |
L5178Y (Lymphoma Cells) (cluster #1 Of 2), Other |
Other |
2 |
0.72 |
Functional ≤ 10μM
|
|
Z80268-2-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80269-3-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #3 Of 3), Other |
Other |
3500 |
0.45 |
Functional ≤ 10μM |
|
Z80347-2-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
15 |
0.64 |
Functional ≤ 10μM |
|
Z80525-2-O |
SW48 (cluster #2 Of 2), Other |
Other |
130 |
0.57 |
Functional ≤ 10μM |
|
Z80532-1-O |
T-24 (Bladder Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80682-9-O |
A549 (Lung Carcinoma Cells) (cluster #9 Of 11), Other |
Other |
1000 |
0.49 |
Functional ≤ 10μM |
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
50 |
0.60 |
Functional ≤ 10μM |
|
Z81048-1-O |
TSU (Prostatic Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
58 |
0.60 |
Functional ≤ 10μM
|
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
760 |
0.50 |
Functional ≤ 10μM |
|
Z81137-1-O |
L929 (Fibroblast Cells) (cluster #1 Of 2), Other |
Other |
7700 |
0.42 |
Functional ≤ 10μM
|
|
Z81170-4-O |
LNCaP (Prostate Carcinoma) (cluster #4 Of 5), Other |
Other |
69 |
0.59 |
Functional ≤ 10μM
|
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
3 |
0.70 |
Functional ≤ 10μM |
|
Z81335-3-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
49 |
0.60 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.72 |
-4.03 |
-17.41 |
3 |
7 |
0 |
105 |
246.194 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.26 |
-6.78 |
-57.51 |
2 |
7 |
-1 |
108 |
245.186 |
2 |
↓
|
|
|
|
Analogs
-
3977743
-

-
3977744
-

-
4045293
-

-
1457
-

Draw
Identity
99%
90%
80%
70%
Vendors
And 10 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYSY-1-E |
Thymidylate Synthase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
7300 |
0.42 |
Binding ≤ 10μM
|
|
Z80064-6-O |
CCRF-CEM (T-cell Leukemia) (cluster #6 Of 9), Other |
Other |
500 |
0.52 |
Functional ≤ 10μM
|
|
Z80101-2-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
650 |
0.51 |
Functional ≤ 10μM |
|
Z80120-3-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
92 |
0.58 |
Functional ≤ 10μM |
|
Z80164-4-O |
HT-1080 (Fibrosarcoma Cells) (cluster #4 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
45 |
0.61 |
Functional ≤ 10μM
|
|
Z80208-1-O |
L5178Y (Lymphoma Cells) (cluster #1 Of 2), Other |
Other |
2 |
0.72 |
Functional ≤ 10μM
|
|
Z80268-2-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80269-3-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #3 Of 3), Other |
Other |
3500 |
0.45 |
Functional ≤ 10μM |
|
Z80347-2-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
15 |
0.64 |
Functional ≤ 10μM |
|
Z80525-2-O |
SW48 (cluster #2 Of 2), Other |
Other |
130 |
0.57 |
Functional ≤ 10μM |
|
Z80532-1-O |
T-24 (Bladder Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80682-9-O |
A549 (Lung Carcinoma Cells) (cluster #9 Of 11), Other |
Other |
1000 |
0.49 |
Functional ≤ 10μM |
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
50 |
0.60 |
Functional ≤ 10μM |
|
Z81048-1-O |
TSU (Prostatic Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
58 |
0.60 |
Functional ≤ 10μM
|
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
760 |
0.50 |
Functional ≤ 10μM |
|
Z81137-1-O |
L929 (Fibroblast Cells) (cluster #1 Of 2), Other |
Other |
7700 |
0.42 |
Functional ≤ 10μM
|
|
Z81170-4-O |
LNCaP (Prostate Carcinoma) (cluster #4 Of 5), Other |
Other |
69 |
0.59 |
Functional ≤ 10μM
|
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
3 |
0.70 |
Functional ≤ 10μM |
|
Z81335-3-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
49 |
0.60 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.72 |
-4.23 |
-22.26 |
3 |
7 |
0 |
105 |
246.194 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.26 |
-6.98 |
-63.95 |
2 |
7 |
-1 |
108 |
245.186 |
2 |
↓
|
|
|
|
Analogs
-
3977743
-

-
3977744
-

-
4045293
-

-
1457
-

Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYSY-1-E |
Thymidylate Synthase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
7300 |
0.42 |
Binding ≤ 10μM
|
|
Z80064-6-O |
CCRF-CEM (T-cell Leukemia) (cluster #6 Of 9), Other |
Other |
500 |
0.52 |
Functional ≤ 10μM
|
|
Z80101-2-O |
COLO 320DM (Colon Adenocarcinoma Cells) (cluster #2 Of 2), Other |
Other |
650 |
0.51 |
Functional ≤ 10μM |
|
Z80120-3-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
92 |
0.58 |
Functional ≤ 10μM |
|
Z80164-4-O |
HT-1080 (Fibrosarcoma Cells) (cluster #4 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
45 |
0.61 |
Functional ≤ 10μM
|
|
Z80208-1-O |
L5178Y (Lymphoma Cells) (cluster #1 Of 2), Other |
Other |
2 |
0.72 |
Functional ≤ 10μM
|
|
Z80268-2-O |
MKN-28 (Gastric Epithelial Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80269-3-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #3 Of 3), Other |
Other |
3500 |
0.45 |
Functional ≤ 10μM |
|
Z80347-2-O |
NUGC-3 (Gastric Carcinoma Cells) (cluster #2 Of 2), Other |
Other |
15 |
0.64 |
Functional ≤ 10μM |
|
Z80525-2-O |
SW48 (cluster #2 Of 2), Other |
Other |
130 |
0.57 |
Functional ≤ 10μM |
|
Z80532-1-O |
T-24 (Bladder Carcinoma Cells) (cluster #1 Of 6), Other |
Other |
120 |
0.57 |
Functional ≤ 10μM |
|
Z80682-9-O |
A549 (Lung Carcinoma Cells) (cluster #9 Of 11), Other |
Other |
1000 |
0.49 |
Functional ≤ 10μM |
|
Z80686-1-O |
AZ-521 Cell Line (cluster #1 Of 1), Other |
Other |
50 |
0.60 |
Functional ≤ 10μM |
|
Z81048-1-O |
TSU (Prostatic Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
58 |
0.60 |
Functional ≤ 10μM
|
|
Z81121-1-O |
KKLS Cell Line (cluster #1 Of 1), Other |
Other |
760 |
0.50 |
Functional ≤ 10μM |
|
Z81137-1-O |
L929 (Fibroblast Cells) (cluster #1 Of 2), Other |
Other |
7700 |
0.42 |
Functional ≤ 10μM
|
|
Z81170-4-O |
LNCaP (Prostate Carcinoma) (cluster #4 Of 5), Other |
Other |
69 |
0.59 |
Functional ≤ 10μM
|
|
Z81330-1-O |
SNU- (cluster #1 Of 1), Other |
Other |
3 |
0.70 |
Functional ≤ 10μM |
|
Z81335-3-O |
HCT-15 (Colon Adenocarcinoma Cells) (cluster #3 Of 5), Other |
Other |
49 |
0.60 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.72 |
-4.89 |
-10.74 |
3 |
7 |
0 |
105 |
246.194 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.26 |
-7.64 |
-47.75 |
2 |
7 |
-1 |
108 |
245.186 |
2 |
↓
|
|
|
|
Analogs
-
5298929
-
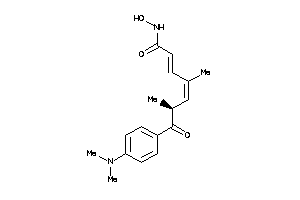
Draw
Identity
99%
90%
80%
70%
Vendors
And 13 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
HDAH-1-B |
Histone Deacetylase-like Amidohydrolase (cluster #1 Of 2), Bacterial |
Bacteria |
700 |
0.39 |
Binding ≤ 10μM
|
|
B1WBY8-1-E |
Histone Deacetylase 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
26 |
0.48 |
Binding ≤ 10μM
|
|
HDA10-3-E |
Histone Deacetylase 10 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
8 |
0.52 |
Binding ≤ 10μM
|
|
HDA11-1-E |
Histone Deacetylase 11 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
7 |
0.52 |
Binding ≤ 10μM
|
|
HDAC1-1-E |
Histone Deacetylase 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
26 |
0.48 |
Binding ≤ 10μM
|
|
HDAC2-2-E |
Histone Deacetylase 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
7600 |
0.33 |
Binding ≤ 10μM
|
|
HDAC3-2-E |
Histone Deacetylase 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
26 |
0.48 |
Binding ≤ 10μM
|
|
HDAC4-3-E |
Histone Deacetylase 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
26 |
0.48 |
Binding ≤ 10μM
|
|
HDAC5-3-E |
Histone Deacetylase 5 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
38 |
0.47 |
Binding ≤ 10μM
|
|
HDAC6-2-E |
Histone Deacetylase 6 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
81 |
0.45 |
Binding ≤ 10μM
|
|
HDAC7-3-E |
Histone Deacetylase 7 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
26 |
0.48 |
Binding ≤ 10μM
|
|
HDAC8-2-E |
Histone Deacetylase 8 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
960 |
0.38 |
Binding ≤ 10μM |
|
HDAC9-3-E |
Histone Deacetylase 9 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
800 |
0.39 |
Binding ≤ 10μM
|
|
NCOR2-2-E |
Nuclear Receptor Corepressor 2 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
9 |
0.51 |
Binding ≤ 10μM
|
|
Q5RJZ2-1-E |
Histone Deacetylase 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
26 |
0.48 |
Binding ≤ 10μM
|
|
Q94F81-1-E |
Histone Deacetylase HD2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
7 |
0.52 |
Binding ≤ 10μM
|
|
Q99P97-1-E |
Histone Deacetylase 6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
8600 |
0.32 |
Binding ≤ 10μM
|
|
Q9XYC7-1-E |
Histone Deacetylase (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1 |
0.57 |
Binding ≤ 10μM
|
|
Q9ZTP8-1-E |
Histone Deacetylase HD1B (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
SF3B3-1-E |
Splicing Factor 3B Subunit 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
611 |
0.40 |
Binding ≤ 10μM
|
|
TF-1-E |
Coagulation Factor III (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
50 |
0.46 |
Binding ≤ 10μM
|
|
HDA10-1-E |
Histone Deacetylase 10 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDA11-1-E |
Histone Deacetylase 11 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDAC1-1-E |
Histone Deacetylase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1000 |
0.38 |
Functional ≤ 10μM
|
|
HDAC2-1-E |
Histone Deacetylase 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDAC3-1-E |
Histone Deacetylase 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDAC4-1-E |
Histone Deacetylase 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
4000 |
0.34 |
Functional ≤ 10μM
|
|
HDAC5-1-E |
Histone Deacetylase 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDAC6-1-E |
Histone Deacetylase 6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDAC7-1-E |
Histone Deacetylase 7 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDAC8-1-E |
Histone Deacetylase 8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
HDAC9-1-E |
Histone Deacetylase 9 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
100 |
0.45 |
Functional ≤ 10μM
|
|
Z103359-1-O |
NCI-H661 (cluster #1 Of 1), Other |
Other |
100 |
0.45 |
Functional ≤ 10μM
|
|
Z50425-3-O |
Plasmodium Falciparum (cluster #3 Of 22), Other |
Other |
10 |
0.51 |
Functional ≤ 10μM
|
|
Z80123-1-O |
DO4 (Melanoma Cells) (cluster #1 Of 1), Other |
Other |
90 |
0.45 |
Functional ≤ 10μM
|
|
Z80125-1-O |
DU-145 (Prostate Carcinoma) (cluster #1 Of 9), Other |
Other |
60 |
0.46 |
Functional ≤ 10μM
|
|
Z80224-1-O |
MCF7 (Breast Carcinoma Cells) (cluster #1 Of 14), Other |
Other |
20 |
0.49 |
Functional ≤ 10μM
|
|
Z80269-1-O |
MKN-45 (Gastric Adenocarcinoma Cells) (cluster #1 Of 3), Other |
Other |
100 |
0.45 |
Functional ≤ 10μM
|
|
Z80277-1-O |
MM96L (Melanoma Cells) (cluster #1 Of 1), Other |
Other |
30 |
0.48 |
Functional ≤ 10μM
|
|
Z80475-2-O |
SK-BR-3 (Breast Adenocarcinoma) (cluster #2 Of 3), Other |
Other |
20 |
0.49 |
Functional ≤ 10μM
|
|
Z80485-1-O |
SK-MEL-28 (Melanoma Cells) (cluster #1 Of 6), Other |
Other |
280 |
0.42 |
Functional ≤ 10μM
|
|
Z80513-1-O |
SQ20B (cluster #1 Of 1), Other |
Other |
200 |
0.43 |
Functional ≤ 10μM
|
|
Z80525-1-O |
SW48 (cluster #1 Of 2), Other |
Other |
5000 |
0.34 |
Functional ≤ 10μM
|
|
Z80532-3-O |
T-24 (Bladder Carcinoma Cells) (cluster #3 Of 6), Other |
Other |
4000 |
0.34 |
Functional ≤ 10μM
|
|
Z80682-1-O |
A549 (Lung Carcinoma Cells) (cluster #1 Of 11), Other |
Other |
80 |
0.45 |
Functional ≤ 10μM
|
|
Z80725-1-O |
C180-13S (Ovarian Carcinoma Cells) (cluster #1 Of 1), Other |
Other |
60 |
0.46 |
Functional ≤ 10μM
|
|
Z80857-2-O |
Friend Leukemia Cell Line (cluster #2 Of 2), Other |
Other |
40 |
0.47 |
Functional ≤ 10μM
|
|
Z80877-2-O |
NCI-H1299 (Non-small Cell Lung Carcinoma) (cluster #2 Of 2), Other |
Other |
100 |
0.45 |
Functional ≤ 10μM
|
|
Z80928-3-O |
HCT-116 (Colon Carcinoma Cells) (cluster #3 Of 9), Other |
Other |
900 |
0.38 |
Functional ≤ 10μM
|
|
Z80957-1-O |
HMEC (Microvascular Endothelial Cells) (cluster #1 Of 2), Other |
Other |
1600 |
0.37 |
Functional ≤ 10μM
|
|
Z81070-1-O |
JAM Cell Line (cluster #1 Of 1), Other |
Other |
80 |
0.45 |
Functional ≤ 10μM
|
|
Z81199-1-O |
NCI-H446 (Small Cell Lung Carcinoma Cells) (cluster #1 Of 2), Other |
Other |
80 |
0.45 |
Functional ≤ 10μM
|
|
Z81252-3-O |
MDA-MB-231 (Breast Adenocarcinoma Cells) (cluster #3 Of 11), Other |
Other |
90 |
0.45 |
Functional ≤ 10μM
|
|
Z80332-1-O |
NFF (Fibroblast Cells) (cluster #1 Of 1), Other |
Other |
200 |
0.43 |
ADME/T ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.68 |
6.23 |
-20.9 |
2 |
5 |
0 |
70 |
302.374 |
6 |
↓
|
|
Ref
Reference (pH 7)
|
2.68 |
6.23 |
-20.14 |
2 |
5 |
0 |
70 |
302.374 |
6 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.68 |
7.44 |
-64.15 |
1 |
5 |
-1 |
72 |
301.366 |
6 |
↓
|
|