|
|
|
|
|
Analogs
-
13119547
-
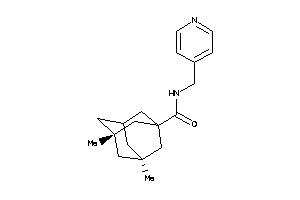
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP17A-2-E |
Cytochrome P450 17A1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
7700 |
0.36 |
Binding ≤ 10μM
|
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
1500 |
0.41 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.80 |
6.79 |
-7.41 |
1 |
3 |
0 |
42 |
270.376 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
2.80 |
7.24 |
-41.4 |
2 |
3 |
1 |
43 |
271.384 |
3 |
↓
|
|
|
|
|
|
|
Analogs
-
120294
-
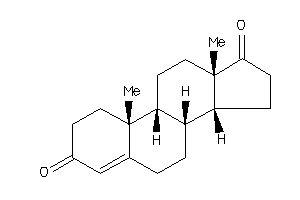
-
2046798
-
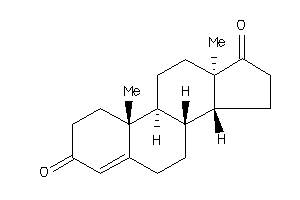
Draw
Identity
99%
90%
80%
70%
Vendors
And 15 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.06 |
9.29 |
-10.68 |
0 |
2 |
0 |
34 |
286.415 |
0 |
↓
|
|
Ref
Reference (pH 7)
|
3.06 |
9.39 |
-9.03 |
0 |
2 |
0 |
34 |
286.415 |
0 |
↓
|
|
|
|
|
|
|
Analogs
-
5275588
-
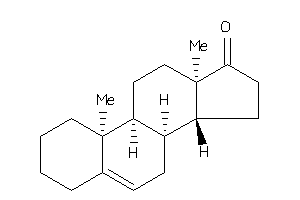
-
5276420
-
-
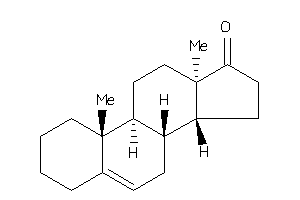
5276422
-
-
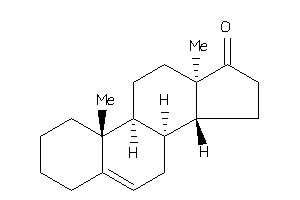
6067588
-
-
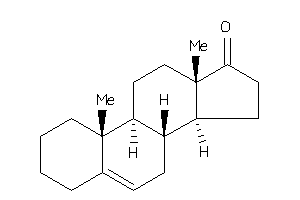
6394326
-
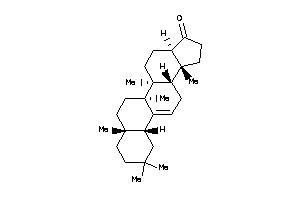
Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-1,2,3,4,7,8,9,11,12,14,15,16-dodecahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,3,4,7,8,9,11,…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
660 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.63 |
2.34 |
-4 |
0 |
1 |
0 |
17 |
272.432 |
0 |
↓
|
|
|
|
Analogs
-
5276420
-

-
5276422
-

-
6067588
-

-
6394326
-

-
39942009
-
Draw
Identity
99%
90%
80%
70%
Popular Name:
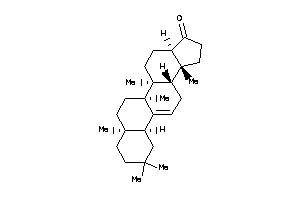
10,13-dimethyl-1,2,3,4,7,8,9,11,12,14,15,16-dodecahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,3,4,7,8,9,11,…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
660 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.63 |
2.31 |
-4.18 |
0 |
1 |
0 |
17 |
272.432 |
0 |
↓
|
|
|
|
Analogs
-
5276422
-

-
6067588
-

-
6394326
-

-
39942009
-

-
39942010
-
Draw
Identity
99%
90%
80%
70%
Popular Name:
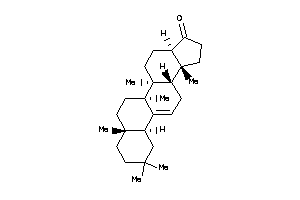
10,13-dimethyl-1,2,3,4,7,8,9,11,12,14,15,16-dodecahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,3,4,7,8,9,11,…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
660 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.63 |
2.31 |
-4.04 |
0 |
1 |
0 |
17 |
272.432 |
0 |
↓
|
|
|
|
Analogs
-
6067588
-

-
6394326
-

-
39942009
-

-
39942010
-

-
5275586
-
Draw
Identity
99%
90%
80%
70%
Popular Name:
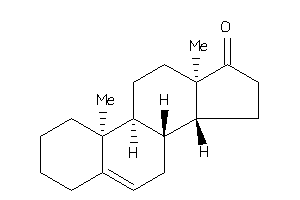
10,13-dimethyl-1,2,3,4,7,8,9,11,12,14,15,16-dodecahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,3,4,7,8,9,11,…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
660 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.63 |
2.21 |
-4.29 |
0 |
1 |
0 |
17 |
272.432 |
0 |
↓
|
|
|
|
Analogs
-
120294
-

-
2046798
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
13-methyl-2,3,6,7,8,9,10,11,12,13,14,15,16,17-tetradecahydro-1H-cyclopenta[a]phenanthrene-3,17-dione
13-methyl-2,3,6,7,8,9,10,11,12,1…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
140 |
0.48 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.82 |
8.94 |
-14.13 |
0 |
2 |
0 |
34 |
272.388 |
0 |
↓
|
|
|
|
Analogs
-
120294
-

-
2046798
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
13-methyl-2,3,6,7,8,9,10,11,12,13,14,15,16,17-tetradecahydro-1H-cyclopenta[a]phenanthrene-3,17-dione
13-methyl-2,3,6,7,8,9,10,11,12,1…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
140 |
0.48 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.82 |
9.31 |
-10.31 |
0 |
2 |
0 |
34 |
272.388 |
0 |
↓
|
|
|
|
Analogs
-
120294
-

-
2046798
-

Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
140 |
0.48 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.82 |
9.03 |
-10.73 |
0 |
2 |
0 |
34 |
272.388 |
0 |
↓
|
|
|
|
Analogs
-
120294
-

-
2046798
-

Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
140 |
0.48 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.82 |
9.17 |
-10.74 |
0 |
2 |
0 |
34 |
272.388 |
0 |
↓
|
|
|
|
Analogs
-
4074121
-
-
4074122
-
-
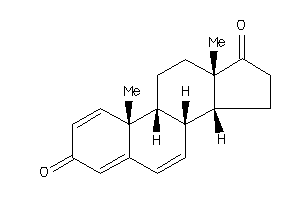
4074123
-
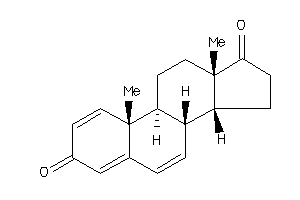
-
35780670
-
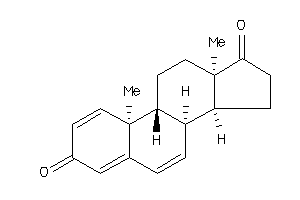
-
35780673
-
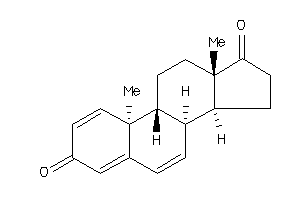
Draw
Identity
99%
90%
80%
70%
Popular Name:
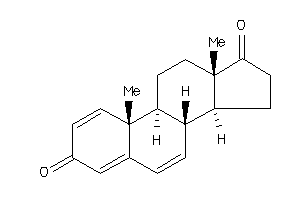
10,13-dimethyl-9,11,12,14,15,16-hexahydro-8H-cyclopenta[a]phenanthrene-3,17-dione
10,13-dimethyl-9,11,12,14,15,16-…
Find On:
PubMed —
Wikipedia —
Google
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
180 |
0.45 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.53 |
9.36 |
-15.34 |
0 |
2 |
0 |
34 |
282.383 |
0 |
↓
|
|
|
|
Analogs
-
4074122
-

-
4074123
-

-
35780670
-

-
35780673
-

-
35780674
-
Draw
Identity
99%
90%
80%
70%
Popular Name:
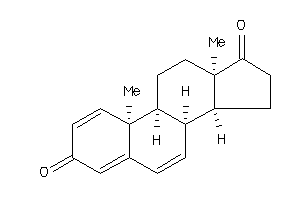
1,4,6-Andorstatriene-3,17-dione; 1,4,6-Androstatriene-3,17-dione; 1,4,6-androstrien-3,17-dione; A146-T317D; ATD; Androsta-1,4,6-triene-3,17-dione; C19H22O2; LS-19416
1,4,6-Andorstatriene-3,17-dione;…
Find On:
PubMed —
Wikipedia —
Google
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
180 |
0.45 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.53 |
9.2 |
-13.21 |
0 |
2 |
0 |
34 |
282.383 |
0 |
↓
|
|
|
|
|
|
|
Analogs
-
1041161
-
-
3830799
-
Draw
Identity
99%
90%
80%
70%
Popular Name:
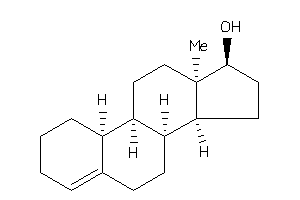
10,13-dimethyl-2,3,6,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,6,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
830 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
-0.27 |
-1.82 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
-
1041161
-

-
3830799
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-2,3,6,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,6,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
830 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
0.12 |
-1.53 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
-
1041161
-

-
3830799
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-2,3,6,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,6,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
830 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
-0.19 |
-2.01 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
-
1041161
-

-
3830799
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-2,3,6,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,6,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
830 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
-0.15 |
-1.87 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
15 |
0.61 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.45 |
8.02 |
-12.62 |
0 |
3 |
0 |
43 |
244.29 |
0 |
↓
|
|
|
|
Analogs
-
5275931
-
-
5276670
-
-
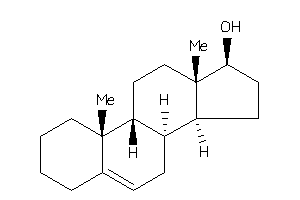
5276759
-
-
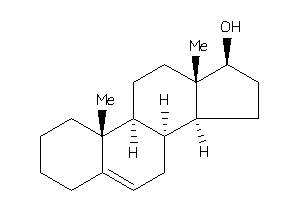
4215047
-
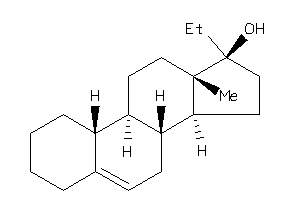
Draw
Identity
99%
90%
80%
70%
Popular Name:
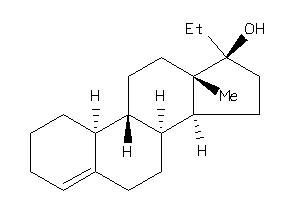
10,13-dimethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,4,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
-0.17 |
-1.8 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
-
5276670
-

-
5276759
-

-
5275835
-
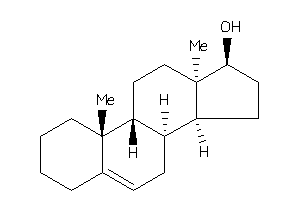
-
4215047
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,4,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
-0.04 |
-1.52 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
-
5276759
-

-
5275835
-

-
5275931
-

-
4215047
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
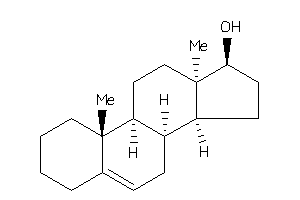
10,13-dimethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,4,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
-0.26 |
-1.78 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
-
5275835
-

-
5275931
-

-
5276670
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-17-ol
10,13-dimethyl-2,3,4,7,8,9,11,12…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.82 |
-0.22 |
-1.71 |
1 |
1 |
0 |
20 |
274.448 |
0 |
↓
|
|
|
|
Analogs
-
5275910
-
-
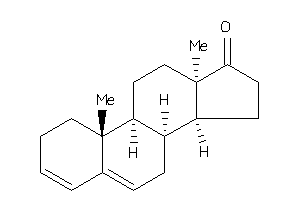
5276648
-
-
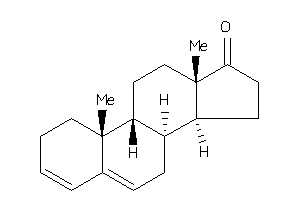
5276737
-
-
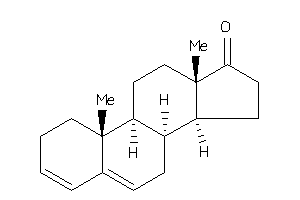
17002216
-
-
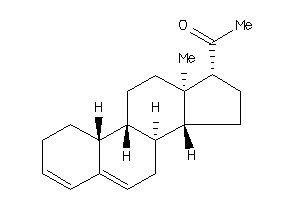
17002220
-
Draw
Identity
99%
90%
80%
70%
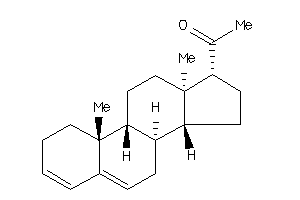
Popular Name:
10,13-dimethyl-1,2,7,8,9,11,12,14,15,16-decahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,7,8,9,11,12,1…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
58 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.38 |
2.31 |
-4.54 |
0 |
1 |
0 |
17 |
270.416 |
0 |
↓
|
|
|
|
Analogs
-
5276648
-

-
5276737
-

-
17002216
-

-
17002220
-

-
5275813
-
Draw
Identity
99%
90%
80%
70%
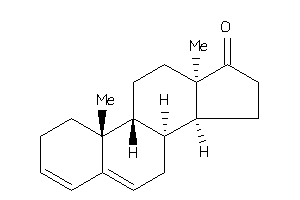
Popular Name:
10,13-dimethyl-1,2,7,8,9,11,12,14,15,16-decahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,7,8,9,11,12,1…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
58 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.38 |
2.09 |
-5.81 |
0 |
1 |
0 |
17 |
270.416 |
0 |
↓
|
|
|
|
Analogs
-
5276737
-

-
17002216
-

-
17002220
-

-
5275813
-

-
5275910
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-1,2,7,8,9,11,12,14,15,16-decahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,7,8,9,11,12,1…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
58 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.38 |
2.38 |
-4.94 |
0 |
1 |
0 |
17 |
270.416 |
0 |
↓
|
|
|
|
Analogs
-
17002216
-

-
17002220
-

-
5275813
-

-
5275910
-

Draw
Identity
99%
90%
80%
70%
Popular Name:
10,13-dimethyl-1,2,7,8,9,11,12,14,15,16-decahydrocyclopenta[a]phenanthren-17-one
10,13-dimethyl-1,2,7,8,9,11,12,1…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CP19A-2-E |
Cytochrome P450 19A1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
58 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.38 |
2.31 |
-4.6 |
0 |
1 |
0 |
17 |
270.416 |
0 |
↓
|
|