|
|
Analogs
-
34372551
-
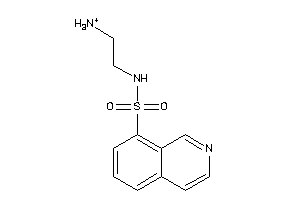
-
41661539
-
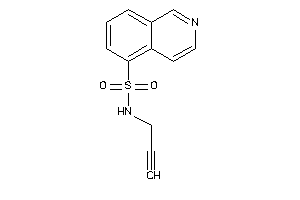
Draw
Identity
99%
90%
80%
70%
Vendors
And 14 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
KAPCA-2-E |
CAMP-dependent Protein Kinase Alpha-catalytic Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
2000 |
0.47 |
Binding ≤ 10μM
|
|
KAPCB-2-E |
CAMP-dependent Protein Kinase Beta-1 Catalytic Subunit (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
2000 |
0.47 |
Binding ≤ 10μM
|
|
KAPCG-1-E |
CAMP-dependent Protein Kinase, Gamma Catalytic Subunit (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2000 |
0.47 |
Binding ≤ 10μM
|
|
KPCA-6-E |
Protein Kinase C Alpha (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCB-3-E |
Protein Kinase C Beta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCD-4-E |
Protein Kinase C Delta (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCD1-2-E |
Protein Kinase C Mu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCD3-2-E |
Protein Kinase C Nu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCE-4-E |
Protein Kinase C Epsilon (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCG-2-E |
Protein Kinase C Gamma (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCI-2-E |
Protein Kinase C Iota (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCL-3-E |
Protein Kinase C Eta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCT-1-E |
Protein Kinase C Theta (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
KPCZ-4-E |
Protein Kinase C Zeta (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
7000 |
0.42 |
Binding ≤ 10μM
|
|
P90584-2-E |
Protein Kinase Pfmrk (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
700 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.12 |
-0.55 |
-55.51 |
4 |
5 |
1 |
87 |
252.319 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
-0.12 |
-0.94 |
-12.15 |
3 |
5 |
0 |
85 |
251.311 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
KAPCA-2-E |
CAMP-dependent Protein Kinase Alpha-catalytic Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
3500 |
0.38 |
Binding ≤ 10μM
|
|
KAPCB-2-E |
CAMP-dependent Protein Kinase Beta-1 Catalytic Subunit (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3500 |
0.38 |
Binding ≤ 10μM
|
|
KAPCG-1-E |
CAMP-dependent Protein Kinase, Gamma Catalytic Subunit (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3500 |
0.38 |
Binding ≤ 10μM
|
|
KPCA-6-E |
Protein Kinase C Alpha (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCB-3-E |
Protein Kinase C Beta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD-4-E |
Protein Kinase C Delta (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD1-2-E |
Protein Kinase C Mu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD3-2-E |
Protein Kinase C Nu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCE-4-E |
Protein Kinase C Epsilon (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCG-2-E |
Protein Kinase C Gamma (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCI-2-E |
Protein Kinase C Iota (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCL-3-E |
Protein Kinase C Eta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCT-1-E |
Protein Kinase C Theta (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCZ-4-E |
Protein Kinase C Zeta (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
KAPCA-2-E |
CAMP-dependent Protein Kinase Alpha-catalytic Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM |
|
KAPCA-2-E |
CAMP-dependent Protein Kinase Alpha-catalytic Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
3500 |
0.38 |
Binding ≤ 10μM
|
|
KAPCB-2-E |
CAMP-dependent Protein Kinase Beta-1 Catalytic Subunit (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM |
|
KAPCB-2-E |
CAMP-dependent Protein Kinase Beta-1 Catalytic Subunit (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3500 |
0.38 |
Binding ≤ 10μM
|
|
KAPCG-1-E |
CAMP-dependent Protein Kinase, Gamma Catalytic Subunit (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM |
|
KAPCG-1-E |
CAMP-dependent Protein Kinase, Gamma Catalytic Subunit (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3500 |
0.38 |
Binding ≤ 10μM
|
|
KGP1-1-E |
CGMP-dependent Protein Kinase 1 Beta (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
5800 |
0.37 |
Binding ≤ 10μM
|
|
KGP2-1-E |
CGMP-dependent Protein Kinase 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
5800 |
0.37 |
Binding ≤ 10μM
|
|
KPCA-6-E |
Protein Kinase C Alpha (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCA-6-E |
Protein Kinase C Alpha (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCB-3-E |
Protein Kinase C Beta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCB-3-E |
Protein Kinase C Beta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD-4-E |
Protein Kinase C Delta (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD-4-E |
Protein Kinase C Delta (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD1-2-E |
Protein Kinase C Mu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD1-2-E |
Protein Kinase C Mu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD3-2-E |
Protein Kinase C Nu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCD3-2-E |
Protein Kinase C Nu (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCE-4-E |
Protein Kinase C Epsilon (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCE-4-E |
Protein Kinase C Epsilon (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCG-2-E |
Protein Kinase C Gamma (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCG-2-E |
Protein Kinase C Gamma (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCI-2-E |
Protein Kinase C Iota (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCI-2-E |
Protein Kinase C Iota (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCL-3-E |
Protein Kinase C Eta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCL-3-E |
Protein Kinase C Eta (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCT-1-E |
Protein Kinase C Theta (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCT-1-E |
Protein Kinase C Theta (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCZ-4-E |
Protein Kinase C Zeta (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
KPCZ-4-E |
Protein Kinase C Zeta (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Binding ≤ 10μM
|
|
Q95J97-1-E |
CAMP-dependent Protein Kinase Alpha-catalytic Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
3000 |
0.39 |
Binding ≤ 10μM
|
|
Z50425-3-O |
Plasmodium Falciparum (cluster #3 Of 22), Other |
Other |
5400 |
0.37 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
KPCA-6-E |
Protein Kinase C Alpha (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
7800 |
0.21 |
Binding ≤ 10μM
|
|
Q7ZJM1-1-V |
Human Immunodeficiency Virus Type 1 Integrase (cluster #1 Of 6), Viral |
Viruses |
5000 |
0.22 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.40 |
1.86 |
-18.45 |
4 |
9 |
0 |
144 |
482.514 |
8 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 5 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CDK2-4-E |
Cyclin-dependent Kinase 2 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
3300 |
0.33 |
Binding ≤ 10μM
|
|
CSK-1-E |
Tyrosine-protein Kinase CSK (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
2100 |
0.35 |
Binding ≤ 10μM
|
|
FER-1-E |
Tyrosine-protein Kinase FER (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1 |
0.55 |
Binding ≤ 10μM
|
|
FGFR1-1-E |
Fibroblast Growth Factor Receptor 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1480 |
0.35 |
Binding ≤ 10μM
|
|
FGFR2-1-E |
Fibroblast Growth Factor Receptor 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
940 |
0.37 |
Binding ≤ 10μM
|
|
FGFR3-1-E |
Fibroblast Growth Factor Receptor 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1480 |
0.35 |
Binding ≤ 10μM
|
|
FGFR4-1-E |
Fibroblast Growth Factor Receptor 4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1480 |
0.35 |
Binding ≤ 10μM
|
|
FYN-1-E |
Tyrosine-protein Kinase FYN (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
500 |
0.38 |
Binding ≤ 10μM
|
|
HCK-1-E |
Tyrosine-protein Kinase HCK (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
7700 |
0.31 |
Binding ≤ 10μM
|
|
IKKB-1-E |
Inhibitor Of Nuclear Factor Kappa B Kinase Beta Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.40 |
Binding ≤ 10μM
|
|
JAK1-1-E |
Tyrosine-protein Kinase JAK1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
15 |
0.48 |
Binding ≤ 10μM
|
|
JAK2-1-E |
Tyrosine-protein Kinase JAK2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1 |
0.55 |
Binding ≤ 10μM
|
|
JAK3-2-E |
Tyrosine-protein Kinase JAK3 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
5 |
0.51 |
Binding ≤ 10μM
|
|
KAP3-1-E |
CAMP-dependent Protein Kinase Type II-beta Regulatory Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
7100 |
0.31 |
Binding ≤ 10μM
|
|
KAPCA-2-E |
CAMP-dependent Protein Kinase Alpha-catalytic Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
7100 |
0.31 |
Binding ≤ 10μM
|
|
KAPCB-2-E |
CAMP-dependent Protein Kinase Beta-1 Catalytic Subunit (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
7100 |
0.31 |
Binding ≤ 10μM
|
|
KAPCG-1-E |
CAMP-dependent Protein Kinase, Gamma Catalytic Subunit (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7100 |
0.31 |
Binding ≤ 10μM
|
|
KPCA-6-E |
Protein Kinase C Alpha (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
1200 |
0.36 |
Binding ≤ 10μM
|
|
MP2K1-1-E |
Dual Specificity Mitogen-activated Protein Kinase Kinase 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
160 |
0.41 |
Binding ≤ 10μM
|
|
MP2K2-1-E |
Dual Specificity Mitogen-activated Protein Kinase Kinase 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
160 |
0.41 |
Binding ≤ 10μM
|
|
PGFRA-1-E |
Platelet-derived Growth Factor Receptor Alpha (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1490 |
0.35 |
Binding ≤ 10μM
|
|
PGFRB-1-E |
Platelet-derived Growth Factor Receptor Beta (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1490 |
0.35 |
Binding ≤ 10μM
|
|
TYK2-1-E |
Tyrosine-protein Kinase TYK2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1 |
0.55 |
Binding ≤ 10μM
|
|
VGFR1-1-E |
Vascular Endothelial Growth Factor Receptor 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1520 |
0.35 |
Binding ≤ 10μM
|
|
VGFR2-1-E |
Vascular Endothelial Growth Factor Receptor 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1400 |
0.36 |
Binding ≤ 10μM
|
|
VGFR3-1-E |
Vascular Endothelial Growth Factor Receptor 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
690 |
0.38 |
Binding ≤ 10μM
|
|
JAK1-1-E |
Tyrosine-protein Kinase JAK1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
52 |
0.44 |
Functional ≤ 10μM
|
|
JAK3-1-E |
Tyrosine-protein Kinase JAK3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
52 |
0.44 |
Functional ≤ 10μM
|
|
Z104300-1-O |
Mitogen-activated Protein Kinase; ERK1/ERK2 (cluster #1 Of 1), Other |
Other |
1780 |
0.35 |
Binding ≤ 10μM
|
|
Z80109-2-O |
CTLL (Cytotoxic T-cells) (cluster #2 Of 2), Other |
Other |
52 |
0.44 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.03 |
7.37 |
-15.5 |
2 |
4 |
0 |
62 |
309.344 |
1 |
↓
|
|