|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
3300 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.46 |
2.04 |
-17.24 |
3 |
4 |
0 |
70 |
203.197 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.46 |
2.98 |
-68.01 |
2 |
4 |
-1 |
72 |
202.189 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 6 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
3600 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.25 |
2.09 |
-16.1 |
3 |
4 |
0 |
70 |
203.197 |
1 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.25 |
3.03 |
-63.83 |
2 |
4 |
-1 |
72 |
202.189 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
6500 |
0.40 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.92 |
4.24 |
-20.3 |
2 |
4 |
0 |
59 |
245.278 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.92 |
5.49 |
-58.63 |
1 |
4 |
-1 |
61 |
244.27 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
6500 |
0.40 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.92 |
4.24 |
-20.32 |
2 |
4 |
0 |
59 |
245.278 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.92 |
5.49 |
-58.67 |
1 |
4 |
-1 |
61 |
244.27 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 15 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
5000 |
0.41 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.22 |
8.63 |
-40.06 |
0 |
2 |
-1 |
40 |
255.68 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
3200 |
0.37 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.73 |
10.42 |
-47.49 |
0 |
2 |
-1 |
40 |
271.295 |
1 |
↓
|
|
Mid
Mid (pH 6-8)
|
4.05 |
3.43 |
-16.08 |
0 |
2 |
0 |
34 |
272.303 |
1 |
↓
|
|
|
|
Analogs
-
5284667
-
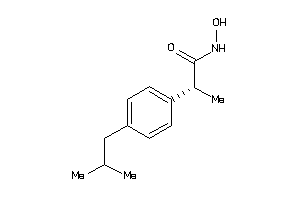
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
5700 |
0.46 |
Binding ≤ 10μM
|
|
PGH1-2-E |
Cyclooxygenase-1 (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
10000 |
0.44 |
Binding ≤ 10μM
|
|
PGH2-3-E |
Cyclooxygenase-2 (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
10000 |
0.44 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.01 |
5.41 |
-13.02 |
2 |
3 |
0 |
49 |
221.3 |
4 |
↓
|
|
Ref
Reference (pH 7)
|
3.19 |
3.38 |
-4.5 |
2 |
3 |
0 |
53 |
221.3 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
3.01 |
6.62 |
-54.94 |
1 |
3 |
-1 |
52 |
220.292 |
4 |
↓
|
|
|
|
Analogs
-
381
-
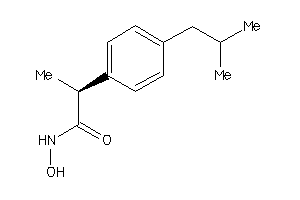
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
5700 |
0.46 |
Binding ≤ 10μM
|
|
PGH1-2-E |
Cyclooxygenase-1 (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
10000 |
0.44 |
Binding ≤ 10μM
|
|
PGH2-3-E |
Cyclooxygenase-2 (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
10000 |
0.44 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.01 |
5.4 |
-12.87 |
2 |
3 |
0 |
49 |
221.3 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
3.01 |
6.62 |
-54.37 |
1 |
3 |
-1 |
52 |
220.292 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
50 |
0.49 |
Binding ≤ 10μM
|
|
LX12L-1-E |
Arachidonate 12-lipoxygenase (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.84 |
0.32 |
-8.91 |
1 |
3 |
0 |
40 |
283.371 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
3.84 |
0.77 |
-44.67 |
0 |
3 |
-1 |
43 |
282.363 |
7 |
↓
|
|
|
|
Analogs
-
3947435
-
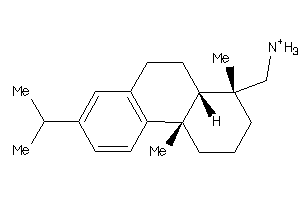
-
3947436
-
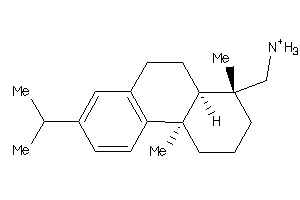
-
3947437
-
Draw
Identity
99%
90%
80%
70%
Vendors
And 10 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
8000 |
0.34 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.23 |
9.43 |
-47.71 |
3 |
1 |
1 |
28 |
286.483 |
2 |
↓
|
|
|
|
Analogs
-
3947435
-

-
3947436
-

-
3947437
-

Draw
Identity
99%
90%
80%
70%
Vendors
And 30 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
8000 |
0.34 |
Binding ≤ 10μM |
|
PDK1-1-E |
Pyruvate Dehydrogenase Kinase Isoform 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
9500 |
0.33 |
Binding ≤ 10μM
|
|
PDK2-1-E |
Pyruvate Dehydrogenase Kinase Isoform 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
9500 |
0.33 |
Binding ≤ 10μM
|
|
PDK3-1-E |
Pyruvate Dehydrogenase Kinase Isoform 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
9500 |
0.33 |
Binding ≤ 10μM
|
|
PDK4-1-E |
Pyruvate Dehydrogenase Kinase Isoform 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
9500 |
0.33 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.23 |
9.42 |
-46.88 |
3 |
1 |
1 |
28 |
286.483 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
460 |
0.44 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.44 |
-0.12 |
-10.39 |
1 |
4 |
0 |
49 |
271.316 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
PGH1-2-E |
Cyclooxygenase-1 (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
3400 |
0.43 |
Binding ≤ 10μM
|
|
PGH2-2-E |
Cyclooxygenase-2 (cluster #2 Of 8), Eukaryotic |
Eukaryotes |
3400 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.79 |
0.32 |
-6.17 |
1 |
1 |
0 |
20 |
234.298 |
2 |
↓
|
|
|
|
|
|
|
Analogs
-
5362954
-
-
17424459
-
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
9500 |
0.47 |
Binding ≤ 10μM
|
|
PGH2-1-E |
Cyclooxygenase-2 (cluster #1 Of 8), Eukaryotic |
Eukaryotes |
9020 |
0.47 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.53 |
1.42 |
-6.86 |
0 |
1 |
0 |
17 |
214.289 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
3000 |
0.30 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.00 |
0.31 |
-46.7 |
2 |
4 |
1 |
42 |
349.454 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 29 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.01 |
6.6 |
-5.41 |
2 |
2 |
0 |
40 |
266.34 |
5 |
↓
|
|
|
|
Analogs
-
6093906
-
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
500 |
0.34 |
Binding ≤ 10μM |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
70 |
0.39 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.19 |
0.43 |
-12.26 |
0 |
6 |
0 |
63 |
348.31 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.47 |
7.18 |
-11.04 |
1 |
4 |
0 |
50 |
283.327 |
5 |
↓
|
|
Hi
High (pH 8-9.5)
|
3.47 |
8.14 |
-43.86 |
0 |
4 |
-1 |
53 |
282.319 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
290 |
0.48 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.56 |
9.49 |
-7.38 |
0 |
2 |
0 |
34 |
248.281 |
2 |
↓
|
|
|
|
Analogs
-
4794131
-
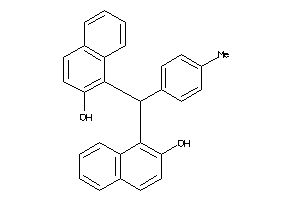
-
4794412
-
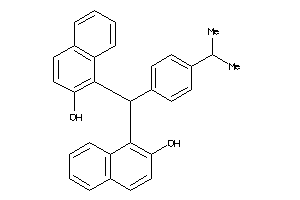
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
3900 |
0.42 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.00 |
0.24 |
-5.57 |
1 |
1 |
0 |
20 |
234.298 |
2 |
↓
|
|
|
|
Analogs
-
39119086
-
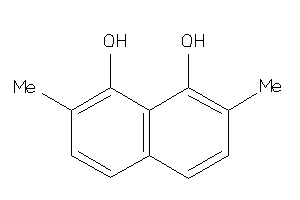
Draw
Identity
99%
90%
80%
70%
Vendors
And 15 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
130 |
0.80 |
Binding ≤ 10μM
|
|
PGH1-2-E |
Cyclooxygenase-1 (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
3000 |
0.64 |
Binding ≤ 10μM
|
|
PGH2-2-E |
Cyclooxygenase-2 (cluster #2 Of 8), Eukaryotic |
Eukaryotes |
3000 |
0.64 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.28 |
4.71 |
-5.14 |
1 |
1 |
0 |
20 |
158.2 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 45 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
3600 |
0.69 |
Binding ≤ 10μM
|
|
PGH1-2-E |
Cyclooxygenase-1 (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
2000 |
0.73 |
Binding ≤ 10μM
|
|
PGH2-2-E |
Cyclooxygenase-2 (cluster #2 Of 8), Eukaryotic |
Eukaryotes |
2000 |
0.73 |
Binding ≤ 10μM
|
|
CP1A2-2-E |
Cytochrome P450 1A2 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3200 |
0.70 |
ADME/T ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.88 |
3.98 |
-5.13 |
1 |
1 |
0 |
20 |
144.173 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
56 |
0.73 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.65 |
6.17 |
-6.72 |
1 |
1 |
0 |
20 |
184.238 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LOX5-6-E |
Arachidonate 5-lipoxygenase (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
130 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.95 |
8.97 |
-8.64 |
1 |
1 |
0 |
20 |
252.288 |
2 |
↓
|
|