|
|
|
|
|
|
|
|
Analogs
-
3995890
-
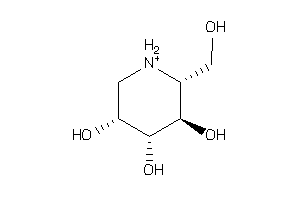
Draw
Identity
99%
90%
80%
70%
Vendors
And 2 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.40 |
-8.74 |
-34.63 |
6 |
5 |
1 |
98 |
164.181 |
1 |
↓
|
|
|
|
|
|
|
Analogs
-
3995890
-

Draw
Identity
99%
90%
80%
70%
Vendors
And 18 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
B1WC34-1-E |
Alpha-glucosidase II Beta Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4600 |
0.68 |
Binding ≤ 10μM
|
|
D3ZAN3-1-E |
Alpha-glucosidase II (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4600 |
0.68 |
Binding ≤ 10μM
|
|
GANAB-1-E |
Neutral Alpha-glucosidase AB (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
160 |
0.86 |
Binding ≤ 10μM |
|
GANC-1-E |
Neutral Alpha-glucosidase C (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.83 |
Binding ≤ 10μM
|
|
GDE-1-E |
Glycogen Debranching Enzyme (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
190 |
0.86 |
Binding ≤ 10μM
|
|
GLCM-2-E |
Glucosylceramidase (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
2000 |
0.73 |
Binding ≤ 10μM |
|
LYAG-1-E |
Alpha-glucosidase (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
400 |
0.81 |
Binding ≤ 10μM
|
|
MGA-1-E |
Maltase-glucoamylase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
40 |
0.94 |
Binding ≤ 10μM
|
|
Q43014-2-E |
Beta-glucosidase (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
9500 |
0.64 |
Binding ≤ 10μM |
|
Q9LGC6-1-E |
Alpha-glucosidase (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
50 |
0.93 |
Binding ≤ 10μM
|
|
SUIS-1-E |
Sucrase-isomaltase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
650 |
0.79 |
Binding ≤ 10μM |
|
P94451-1-B |
Alpha-glucosidase (cluster #1 Of 1), Bacterial |
Bacteria |
1670 |
0.74 |
Binding ≤ 10μM
|
|
Z81057-3-O |
HUVEC (Umbilical Vein Endothelial Cells) (cluster #3 Of 4), Other |
Other |
2000 |
0.73 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.40 |
-9.17 |
-39.08 |
6 |
5 |
1 |
98 |
164.181 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 17 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
LYAG-1-E |
Alpha-glucosidase (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
200 |
0.52 |
Binding ≤ 10μM
|
|
SUIS-1-E |
Sucrase-isomaltase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
70 |
0.56 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.98 |
-11.18 |
-31.61 |
9 |
8 |
1 |
158 |
268.286 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
D3ZJF9-1-E |
Alpha-galactosidase A (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4500 |
0.58 |
Binding ≤ 10μM
|
|
GANC-1-E |
Neutral Alpha-glucosidase C (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
22 |
0.82 |
Binding ≤ 10μM
|
|
GDE-1-E |
Glycogen Debranching Enzyme (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
110 |
0.75 |
Binding ≤ 10μM
|
|
LYAG-1-E |
Alpha-glucosidase (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
8400 |
0.55 |
Binding ≤ 10μM
|
|
MGA-1-E |
Maltase-glucoamylase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
8400 |
0.55 |
Binding ≤ 10μM
|
|
SUIS-1-E |
Sucrase-isomaltase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
700 |
0.66 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.05 |
-12.28 |
-29.67 |
7 |
6 |
1 |
117 |
194.207 |
2 |
↓
|
|
|
|
|
|
|
|