|
|
Analogs
-
32785912
-
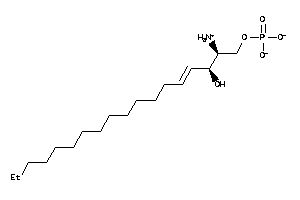
-
36178158
-
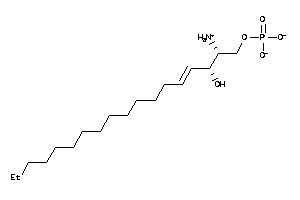
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
6 |
0.46 |
Binding ≤ 10μM
|
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
9 |
0.45 |
Binding ≤ 10μM
|
|
S1PR2-1-E |
Sphingosine 1-phosphate Receptor Edg-5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
S1PR2-1-E |
Sphingosine 1-phosphate Receptor Edg-5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
9 |
0.45 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
2 |
0.49 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
9 |
0.45 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
1 |
0.50 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
9 |
0.45 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
23 |
0.43 |
Binding ≤ 10μM
|
|
S1PR5-2-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
2 |
0.49 |
Binding ≤ 10μM
|
|
S1PR5-2-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
9 |
0.45 |
Binding ≤ 10μM
|
|
S1PR1-3-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Functional ≤ 10μM
|
|
S1PR3-1-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.48 |
-3.29 |
-114.81 |
4 |
6 |
-1 |
120 |
378.47 |
17 |
↓
|
|
|
|
Analogs
-
8860500
-
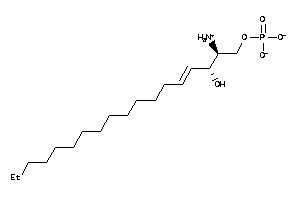
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
6 |
0.46 |
Binding ≤ 10μM
|
|
S1PR2-1-E |
Sphingosine 1-phosphate Receptor Edg-5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
2 |
0.49 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
1 |
0.50 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
23 |
0.43 |
Binding ≤ 10μM
|
|
S1PR5-2-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
2 |
0.49 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.48 |
7.68 |
-113.24 |
4 |
6 |
-1 |
120 |
378.47 |
17 |
↓
|
|
Hi
High (pH 8-9.5)
|
4.48 |
7.4 |
-130.44 |
3 |
6 |
-2 |
119 |
377.462 |
17 |
↓
|
|
Lo
Low (pH 4.5-6)
|
4.48 |
6.53 |
-52.59 |
5 |
6 |
0 |
117 |
379.478 |
17 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
1 |
0.48 |
Binding ≤ 10μM
|
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
1 |
0.48 |
Binding ≤ 10μM
|
|
S1PR2-1-E |
Sphingosine 1-phosphate Receptor Edg-5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1100 |
0.32 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
3 |
0.46 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
6 |
0.44 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
19 |
0.42 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
41 |
0.40 |
Binding ≤ 10μM
|
|
S1PR5-1-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3 |
0.46 |
Binding ≤ 10μM
|
|
S1PR5-1-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
40 |
0.40 |
Binding ≤ 10μM
|
|
S1PR1-3-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
1 |
0.48 |
Functional ≤ 10μM
|
|
S1PR1-3-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
23 |
0.41 |
Functional ≤ 10μM
|
|
S1PR3-1-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
12 |
0.43 |
Functional ≤ 10μM
|
|
S1PR3-1-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
29 |
0.41 |
Functional ≤ 10μM
|
|
S1PR4-1-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4 |
0.45 |
Functional ≤ 10μM
|
|
S1PR4-1-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
80 |
0.38 |
Functional ≤ 10μM
|
|
S1PR5-1-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
5 |
0.45 |
Functional ≤ 10μM
|
|
S1PR5-1-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1988 |
0.31 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.06 |
6.88 |
-118.68 |
4 |
6 |
-1 |
120 |
386.449 |
14 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM |
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
1 |
0.48 |
Binding ≤ 10μM
|
|
S1PR2-1-E |
Sphingosine 1-phosphate Receptor Edg-5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1100 |
0.32 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
6 |
0.44 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
6 |
0.44 |
Binding ≤ 10μM |
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
41 |
0.40 |
Binding ≤ 10μM
|
|
S1PR5-1-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
40 |
0.40 |
Binding ≤ 10μM
|
|
S1PR1-3-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
1 |
0.48 |
Functional ≤ 10μM
|
|
S1PR1-3-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
2 |
0.47 |
Functional ≤ 10μM
|
|
S1PR3-1-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
3 |
0.46 |
Functional ≤ 10μM
|
|
S1PR3-1-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
12 |
0.43 |
Functional ≤ 10μM
|
|
S1PR4-1-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4 |
0.45 |
Functional ≤ 10μM
|
|
S1PR4-1-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
22 |
0.41 |
Functional ≤ 10μM
|
|
S1PR5-1-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Functional ≤ 10μM
|
|
S1PR5-1-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
5 |
0.45 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.06 |
6.89 |
-116.88 |
4 |
6 |
-1 |
120 |
386.449 |
14 |
↓
|
|
|
|
Analogs
-
36178158
-

-
8860500
-

Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
S1PR1-2-E |
Sphingosine 1-phosphate Receptor Edg-1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
6 |
0.46 |
Binding ≤ 10μM
|
|
S1PR3-3-E |
Sphingosine 1-phosphate Receptor Edg-3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
2 |
0.49 |
Binding ≤ 10μM
|
|
S1PR4-2-E |
Sphingosine 1-phosphate Receptor Edg-6 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
1 |
0.50 |
Binding ≤ 10μM
|
|
S1PR5-2-E |
Sphingosine 1-phosphate Receptor Edg-8 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
2 |
0.49 |
Binding ≤ 10μM
|
|
Z80156-3-O |
HL-60 (Promyeloblast Leukemia Cells) (cluster #3 Of 12), Other |
Other |
31 |
0.42 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.48 |
7.35 |
-109.58 |
4 |
6 |
-1 |
120 |
378.47 |
17 |
↓
|
|
Hi
High (pH 8-9.5)
|
4.48 |
6.92 |
-131.55 |
3 |
6 |
-2 |
119 |
377.462 |
17 |
↓
|
|
Lo
Low (pH 4.5-6)
|
4.48 |
6.15 |
-50.94 |
5 |
6 |
0 |
117 |
379.478 |
17 |
↓
|
|