|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
4-[[(2R,3R,4R,6R)-3-hydroxy-2-methyl-6-[[(1R,3S)-3,5,12-trihydroxy-3-(2-hydroxyacetyl)-10-methoxy-6,
4-[[(2R,3R,4R,6R)-3-hydroxy-2-me…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.26 |
3.62 |
-78.54 |
7 |
14 |
0 |
237 |
629.615 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
4-[[(2S,3R,4R,6R)-3-hydroxy-2-methyl-6-[[(1R,3S)-3,5,12-trihydroxy-3-(2-hydroxyacetyl)-10-methoxy-6,
4-[[(2S,3R,4R,6R)-3-hydroxy-2-me…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.26 |
3.15 |
-78.03 |
7 |
14 |
0 |
237 |
629.615 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
4-[[(2R,3S,4R,6R)-3-hydroxy-2-methyl-6-[[(1R,3S)-3,5,12-trihydroxy-3-(2-hydroxyacetyl)-10-methoxy-6,
4-[[(2R,3S,4R,6R)-3-hydroxy-2-me…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.26 |
3.29 |
-76.13 |
7 |
14 |
0 |
237 |
629.615 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
4-[[(2S,3S,4R,6R)-3-hydroxy-2-methyl-6-[[(1R,3S)-3,5,12-trihydroxy-3-(2-hydroxyacetyl)-10-methoxy-6,
4-[[(2S,3S,4R,6R)-3-hydroxy-2-me…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.26 |
2.91 |
-75.72 |
7 |
14 |
0 |
237 |
629.615 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
( -)-(7S,9S)-9-Acetyl-9-amino-7-((2-deoxy-beta-D-erythro-pentopyranosyl)oxy)-7,8,9,10-tetrahydro-6,11-dihydroxy-5,12-naphthacenedione; (+-)-(7S,9S)-9-Acetyl-9-amino-7-((2-deoxy-beta-D-erythro-pentopyranosyl)oxy)-7,8,9,10-tetrahydro-6,11-dihydroxy-5,12-nap
( -)-(7S,9S)-9-Acetyl-9-amino-7-…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.06 |
0.2 |
-71.01 |
7 |
10 |
1 |
178 |
484.481 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.06 |
-0.08 |
-18.43 |
6 |
10 |
0 |
177 |
483.473 |
3 |
↓
|
|
|
|
Analogs
-
22049024
-
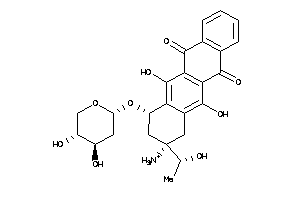
-
22049027
-
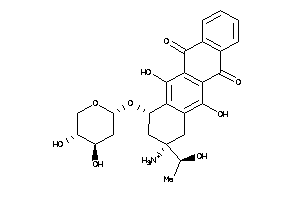
Draw
Identity
99%
90%
80%
70%
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.24 |
-2.28 |
-57.67 |
8 |
10 |
1 |
181 |
486.497 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.24 |
-2.15 |
-14.36 |
7 |
10 |
0 |
180 |
485.489 |
3 |
↓
|
|
|
|
Analogs
-
26171691
-
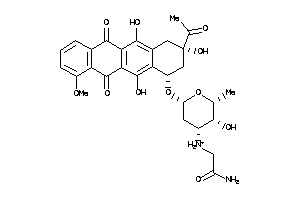
-
43226495
-
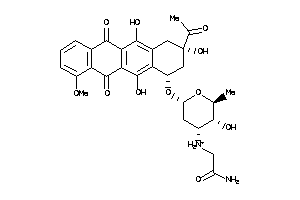
Draw
Identity
99%
90%
80%
70%
Popular Name:
2-[[(2S,3S,4S,6R)-6-[[(1S,3S)-3-acetyl-3,5,12-trihydroxy-10-methoxy-6,11-dioxo-2,4-dihydro-1H-tetrac
2-[[(2S,3S,4S,6R)-6-[[(1S,3S)-3-…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.54 |
0.01 |
-50.69 |
8 |
13 |
1 |
220 |
585.586 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.54 |
-1.15 |
-23.31 |
7 |
13 |
0 |
215 |
584.578 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
N-[(2S,3S,4S,6R)-6-[[(1S,3S)-3-acetyl-3,5,12-trihydroxy-10-methoxy-6,11-dioxo-2,4-dihydro-1H-tetrace
N-[(2S,3S,4S,6R)-6-[[(1S,3S)-3-a…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.17 |
2.39 |
-26.6 |
5 |
12 |
0 |
189 |
569.563 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
2-[[(2S,3S,4S,6R)-6-[[(1S,3S)-3-acetyl-3,5,12-trihydroxy-10-methoxy-6,11-dioxo-2,4-dihydro-1H-tetrac
2-[[(2S,3S,4S,6R)-6-[[(1S,3S)-3-…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.14 |
3.88 |
-47.14 |
6 |
13 |
0 |
217 |
585.562 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.14 |
2.71 |
-66.77 |
5 |
13 |
-1 |
212 |
584.554 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7R,8R,9R)-7-[(2S,4S,5S,6S)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-8-fluoro-6,9,11-trih
(7R,8R,9R)-7-[(2S,4S,5S,6S)-4-am…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.57 |
-1.02 |
-53.47 |
8 |
11 |
1 |
198 |
532.497 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.96 |
0.86 |
-53.82 |
7 |
10 |
1 |
178 |
516.498 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9S)-9-acetyl-7-[(2S,4S,5R,6R)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydr
(7S,9S)-9-acetyl-7-[(2S,4S,5R,6R…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.90 |
0.62 |
-58.2 |
7 |
11 |
1 |
187 |
528.534 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9S)-9-acetyl-7-[(2R,4S,5R,6R)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydr
(7S,9S)-9-acetyl-7-[(2R,4S,5R,6R…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.90 |
1.34 |
-63.72 |
7 |
11 |
1 |
187 |
528.534 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
[(2R,3R,4R,5R,6S)-5-acetoxy-2-[[(1S,3S)-3-acetyl-3,5,12-trihydroxy-6,11-dioxo-2,4-dihydro-1H-tetrace
[(2R,3R,4R,5R,6S)-5-acetoxy-2-[[…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.80 |
8.58 |
-21.71 |
3 |
12 |
0 |
183 |
600.548 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(2S,4S)-4-[(2R,4S,5S,6S)-4-azido-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-2,5,12-trihydroxy-7-met
(2S,4S)-4-[(2R,4S,5S,6S)-4-azido…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.42 |
1.64 |
-52.46 |
4 |
14 |
-1 |
233 |
554.488 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7R,9S)-7-[(2R,4R,5R,6R)-4-(dimethylamino)-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihyd
(7R,9S)-7-[(2R,4R,5R,6R)-4-(dime…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.72 |
3.39 |
-53.08 |
6 |
12 |
1 |
184 |
572.587 |
6 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.72 |
0.96 |
-25.77 |
5 |
12 |
0 |
183 |
571.579 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7R,9S)-7-[(2R,4R,5R,6S)-4-(dimethylamino)-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihyd
(7R,9S)-7-[(2R,4R,5R,6S)-4-(dime…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.72 |
2.92 |
-52.88 |
6 |
12 |
1 |
184 |
572.587 |
6 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.72 |
0.56 |
-26.13 |
5 |
12 |
0 |
183 |
571.579 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7R,9S)-7-[(2R,4R,5S,6R)-4-(dimethylamino)-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihyd
(7R,9S)-7-[(2R,4R,5S,6R)-4-(dime…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.72 |
2.78 |
-51.37 |
6 |
12 |
1 |
184 |
572.587 |
6 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.72 |
0.37 |
-24.7 |
5 |
12 |
0 |
183 |
571.579 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7R,9S)-7-[(2R,4R,5S,6S)-4-(dimethylamino)-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihyd
(7R,9S)-7-[(2R,4R,5S,6S)-4-(dime…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.72 |
2.5 |
-51.54 |
6 |
12 |
1 |
184 |
572.587 |
6 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.72 |
-0.01 |
-24.54 |
5 |
12 |
0 |
183 |
571.579 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7R,9S)-7-[(2R,4R,5S,6R)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydroxy-9-[(E
(7R,9S)-7-[(2R,4R,5S,6R)-4-amino…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.34 |
-0.77 |
-56.65 |
8 |
12 |
1 |
203 |
543.549 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.34 |
-1.17 |
-20.22 |
7 |
12 |
0 |
201 |
542.541 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7R,9S)-7-[(2R,4R,5S,6S)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydroxy-9-[(E
(7R,9S)-7-[(2R,4R,5S,6S)-4-amino…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.34 |
-1.15 |
-56.24 |
8 |
12 |
1 |
203 |
543.549 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.34 |
-1.56 |
-19.92 |
7 |
12 |
0 |
201 |
542.541 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.62 |
1.68 |
-53.37 |
7 |
9 |
1 |
161 |
470.498 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.62 |
1.44 |
-52.93 |
7 |
9 |
1 |
161 |
470.498 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.62 |
1.41 |
-49.59 |
7 |
9 |
1 |
161 |
470.498 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.62 |
1.59 |
-49.58 |
7 |
9 |
1 |
161 |
470.498 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9S)-7-[(2R,4R,5R,6R)-4-amino-5-methoxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydroxy-9-(hy
(7S,9S)-7-[(2R,4R,5R,6R)-4-amino…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.33 |
1.18 |
-54.65 |
7 |
10 |
1 |
170 |
500.524 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.33 |
0.9 |
-14.8 |
6 |
10 |
0 |
169 |
499.516 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9S)-7-[(2R,4R,5R,6S)-4-amino-5-methoxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydroxy-9-(hy
(7S,9S)-7-[(2R,4R,5R,6S)-4-amino…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.33 |
0.84 |
-52.86 |
7 |
10 |
1 |
170 |
500.524 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.33 |
0.54 |
-14.93 |
6 |
10 |
0 |
169 |
499.516 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9S)-7-[(2R,4R,5S,6R)-4-amino-5-methoxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydroxy-9-(hy
(7S,9S)-7-[(2R,4R,5S,6R)-4-amino…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.33 |
0.97 |
-51.39 |
7 |
10 |
1 |
170 |
500.524 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.33 |
0.66 |
-13.44 |
6 |
10 |
0 |
169 |
499.516 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9S)-7-[(2R,4R,5S,6S)-4-amino-5-methoxy-6-methyl-tetrahydropyran-2-yl]oxy-6,9,11-trihydroxy-9-(hy
(7S,9S)-7-[(2R,4R,5S,6S)-4-amino…
Find On:
PubMed —
Wikipedia —
Google
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.33 |
0.6 |
-49.9 |
7 |
10 |
1 |
170 |
500.524 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.33 |
0.2 |
-13.84 |
6 |
10 |
0 |
169 |
499.516 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9R,10R)-10-[(2R,4S,5S,6S)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-9-ethyl-4,6,7,9,11
(7S,9R,10R)-10-[(2R,4S,5S,6S)-4-…
Find On:
PubMed —
Wikipedia —
Google
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z80166-1-O |
HT-29 (Colon Adenocarcinoma Cells) (cluster #1 Of 12), Other |
Other |
1 |
0.34 |
Functional ≤ 10μM
|
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
0 |
0.00 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.96 |
-1.78 |
-56.5 |
9 |
11 |
1 |
202 |
516.523 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.96 |
-2.19 |
-16.68 |
8 |
11 |
0 |
200 |
515.515 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9R,10R)-10-[(2S,4S,5S,6S)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-9-ethyl-4,6,7,9,11
(7S,9R,10R)-10-[(2S,4S,5S,6S)-4-…
Find On:
PubMed —
Wikipedia —
Google
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z80166-1-O |
HT-29 (Colon Adenocarcinoma Cells) (cluster #1 Of 12), Other |
Other |
1 |
0.34 |
Functional ≤ 10μM
|
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
0 |
0.00 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.96 |
-2.08 |
-52.16 |
9 |
11 |
1 |
202 |
516.523 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.96 |
-2.5 |
-14.07 |
8 |
11 |
0 |
200 |
515.515 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9R,10R)-10-[(2R,4R,5S,6S)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-9-ethyl-4,6,7,9,11
(7S,9R,10R)-10-[(2R,4R,5S,6S)-4-…
Find On:
PubMed —
Wikipedia —
Google
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z80166-1-O |
HT-29 (Colon Adenocarcinoma Cells) (cluster #1 Of 12), Other |
Other |
1 |
0.34 |
Functional ≤ 10μM
|
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
0 |
0.00 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.96 |
-2.08 |
-53.98 |
9 |
11 |
1 |
202 |
516.523 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.96 |
-2.08 |
-15.28 |
8 |
11 |
0 |
200 |
515.515 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(7S,9R,10R)-10-[(2S,4R,5S,6S)-4-amino-5-hydroxy-6-methyl-tetrahydropyran-2-yl]oxy-9-ethyl-4,6,7,9,11
(7S,9R,10R)-10-[(2S,4R,5S,6S)-4-…
Find On:
PubMed —
Wikipedia —
Google
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z80166-1-O |
HT-29 (Colon Adenocarcinoma Cells) (cluster #1 Of 12), Other |
Other |
1 |
0.34 |
Functional ≤ 10μM
|
|
Z80193-2-O |
L1210 (Lymphocytic Leukemia Cells) (cluster #2 Of 12), Other |
Other |
0 |
0.00 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.96 |
-2.62 |
-50.7 |
9 |
11 |
1 |
202 |
516.523 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.96 |
-3.02 |
-13.84 |
8 |
11 |
0 |
200 |
515.515 |
3 |
↓
|
|