|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
60 |
0.78 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
400 |
0.69 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.52 |
1.26 |
-72.12 |
6 |
5 |
1 |
105 |
206.291 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.65 |
0.87 |
-52.03 |
5 |
5 |
0 |
106 |
205.283 |
6 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.52 |
0.93 |
-52.68 |
5 |
5 |
0 |
104 |
205.283 |
7 |
↓
|
|
|
|
Analogs
-
39293713
-
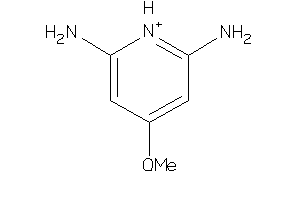
Draw
Identity
99%
90%
80%
70%
Vendors
And 59 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
1000 |
0.93 |
Binding ≤ 10μM
|
|
NOS2-8-E |
Nitric Oxide Synthase, Inducible (cluster #8 Of 9), Eukaryotic |
Eukaryotes |
1000 |
0.93 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
1000 |
0.93 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.53 |
0.78 |
-5.41 |
2 |
3 |
0 |
48 |
124.143 |
1 |
↓
|
|
Mid
Mid (pH 6-8)
|
0.53 |
1.08 |
-29.41 |
3 |
3 |
1 |
49 |
125.151 |
1 |
↓
|
|
|
|
Analogs
-
4556609
-
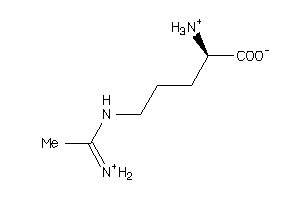
Draw
Identity
99%
90%
80%
70%
Vendors
And 5 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
9000 |
0.59 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
2200 |
0.66 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
4000 |
0.63 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
810 |
0.71 |
Binding ≤ 10μM |
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
1420 |
0.68 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.65 |
0.77 |
-74.95 |
6 |
5 |
1 |
105 |
174.224 |
6 |
↓
|
|
|
|
Analogs
-
19594743
-
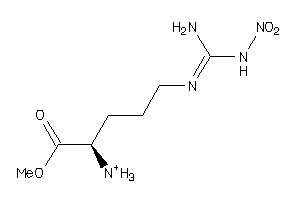
Draw
Identity
99%
90%
80%
70%
Vendors
And 41 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
690 |
0.54 |
Binding ≤ 10μM |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
1600 |
0.51 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
830 |
0.53 |
Binding ≤ 10μM |
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
8000 |
0.45 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
680 |
0.54 |
Binding ≤ 10μM |
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
2700 |
0.49 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
150 |
0.60 |
Functional ≤ 10μM
|
|
NOS3-1-E |
Nitric-oxide Synthase, Endothelial (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2700 |
0.49 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.56 |
1.67 |
-17.55 |
5 |
9 |
0 |
149 |
233.228 |
8 |
↓
|
|
Ref
Reference (pH 7)
|
-1.56 |
1.66 |
-19.79 |
5 |
9 |
0 |
149 |
233.228 |
8 |
↓
|
|
Mid
Mid (pH 6-8)
|
-1.56 |
1.79 |
-39.25 |
4 |
9 |
-1 |
151 |
232.22 |
8 |
↓
|
|
|
|
Analogs
-
44672248
-
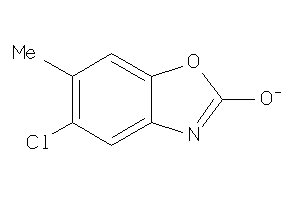
Draw
Identity
99%
90%
80%
70%
Vendors
And 24 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS3-4-E |
Nitric Oxide Synthase, Endothelial (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
8700 |
0.64 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.29 |
-0.12 |
-42.6 |
0 |
3 |
-1 |
49 |
168.559 |
0 |
↓
|
|
Mid
Mid (pH 6-8)
|
1.83 |
2.51 |
-6.85 |
1 |
3 |
0 |
46 |
169.567 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS2-2-E |
Nitric Oxide Synthase, Inducible (cluster #2 Of 9), Eukaryotic |
Eukaryotes |
140 |
1.37 |
Binding ≤ 10μM
|
|
NOS3-5-E |
Nitric Oxide Synthase, Endothelial (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
1260 |
1.18 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.21 |
0.71 |
-33.01 |
4 |
2 |
1 |
52 |
123.176 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.21 |
0.73 |
-5.21 |
3 |
2 |
0 |
50 |
122.168 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
3700 |
1.09 |
Binding ≤ 10μM
|
|
NOS2-2-E |
Nitric Oxide Synthase, Inducible (cluster #2 Of 9), Eukaryotic |
Eukaryotes |
1800 |
1.15 |
Binding ≤ 10μM
|
|
NOS3-7-E |
Nitric Oxide Synthase, Endothelial (cluster #7 Of 7), Eukaryotic |
Eukaryotes |
8200 |
1.02 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.12 |
-2.31 |
-28.02 |
3 |
3 |
1 |
47 |
101.129 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 42 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
9600 |
1.17 |
Binding ≤ 10μM
|
|
NOS2-9-E |
Nitric Oxide Synthase, Inducible (cluster #9 Of 9), Eukaryotic |
Eukaryotes |
4100 |
1.26 |
Binding ≤ 10μM
|
|
NOS3-7-E |
Nitric Oxide Synthase, Endothelial (cluster #7 Of 7), Eukaryotic |
Eukaryotes |
4600 |
1.25 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.05 |
0.71 |
-27.28 |
3 |
2 |
1 |
40 |
85.13 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 8 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
4300 |
0.58 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
860 |
0.65 |
Binding ≤ 10μM |
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
4200 |
0.58 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
100 |
0.75 |
Functional ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Functional ≤ 10μM
|
|
NOS3-1-E |
Nitric-oxide Synthase, Endothelial (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Functional ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3000 |
0.59 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.26 |
1.39 |
-72.47 |
7 |
6 |
1 |
117 |
189.239 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
8.62 |
17.49 |
-47.16 |
0 |
0 |
-1 |
0 |
421.61 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
1600 |
0.90 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.39 |
2.84 |
-28.83 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
1600 |
0.90 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.39 |
2.73 |
-28.82 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
1600 |
0.90 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.39 |
2.84 |
-28.8 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
170 |
1.05 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
1600 |
0.90 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.39 |
2.72 |
-28.86 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Popular Name:
(4R,4aR,7aR)-4-methyl-4,4a,5,6,7,7a-hexahydro-3H-cyclopenta[e]pyridin-2-amine
(4R,4aR,7aR)-4-methyl-4,4a,5,6,7…
Find On:
PubMed —
Wikipedia —
Google
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
31 |
0.96 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
9 |
1.02 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
470 |
0.81 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.79 |
3.74 |
-29.29 |
3 |
2 |
1 |
40 |
153.249 |
0 |
↓
|
|
|
|
Analogs
-
4798508
-
Draw
Identity
99%
90%
80%
70%
Vendors
And 61 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
8700 |
1.01 |
Binding ≤ 10μM
|
|
NOS2-8-E |
Nitric Oxide Synthase, Inducible (cluster #8 Of 9), Eukaryotic |
Eukaryotes |
1900 |
1.14 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
2800 |
1.11 |
Binding ≤ 10μM
|
|
P4K2A-1-E |
PI4-kinase Type II (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7100 |
1.03 |
Binding ≤ 10μM
|
|
P4K2B-1-E |
PI4-kinase Type II Beta (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7100 |
1.03 |
Binding ≤ 10μM
|
|
PI4KA-1-E |
PI4-kinase Alpha Subunit (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7100 |
1.03 |
Binding ≤ 10μM
|
|
PI4KB-2-E |
PI4-kinase Beta Subunit (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
7100 |
1.03 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.52 |
1.48 |
-4.52 |
2 |
2 |
0 |
39 |
94.117 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 25 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.66 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
5500 |
0.49 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
7600 |
0.48 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
320 |
0.61 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3000 |
0.52 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.63 |
1.9 |
-54.25 |
6 |
9 |
0 |
164 |
219.201 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.57 |
-53.16 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.57 |
-48.12 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
100 |
0.58 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.58 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.18 |
-1.74 |
-126.95 |
10 |
9 |
2 |
164 |
249.319 |
10 |
↓
|
|
Mid
Mid (pH 6-8)
|
-3.18 |
-2.12 |
-112.39 |
9 |
9 |
1 |
166 |
248.311 |
10 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-3.18 |
-0.36 |
-249.23 |
11 |
9 |
3 |
168 |
250.327 |
10 |
↓
|
|
|
|
Analogs
-
34223952
-
-
39262116
-
Draw
Identity
99%
90%
80%
70%
Vendors
And 60 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
800 |
0.71 |
Binding ≤ 10μM
|
|
NOS2-4-E |
Nitric Oxide Synthase, Inducible (cluster #4 Of 9), Eukaryotic |
Eukaryotes |
5800 |
0.61 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
2500 |
0.65 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.54 |
3.93 |
-11.35 |
1 |
5 |
0 |
75 |
163.136 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
400 |
1.12 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
500 |
1.10 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
600 |
1.09 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.89 |
2.17 |
-28.15 |
3 |
2 |
1 |
40 |
113.184 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
400 |
1.12 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
500 |
1.10 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
600 |
1.09 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.89 |
2.18 |
-28.17 |
3 |
2 |
1 |
40 |
113.184 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
700 |
1.08 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
6000 |
0.91 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
3000 |
0.97 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.80 |
2.09 |
-28.21 |
3 |
2 |
1 |
40 |
113.184 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
700 |
1.08 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
6000 |
0.91 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
3000 |
0.97 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.80 |
2.1 |
-28.22 |
3 |
2 |
1 |
40 |
113.184 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
240 |
1.32 |
Binding ≤ 10μM
|
|
NOS2-9-E |
Nitric Oxide Synthase, Inducible (cluster #9 Of 9), Eukaryotic |
Eukaryotes |
300 |
1.30 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
630 |
1.24 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.68 |
1.28 |
-28.03 |
3 |
2 |
1 |
38 |
99.157 |
0 |
↓
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-1-E |
Nitric-oxide Synthase, Brain (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.50 |
Binding ≤ 10μM
|
|
NOS2-4-E |
Nitric Oxide Synthase, Inducible (cluster #4 Of 9), Eukaryotic |
Eukaryotes |
90 |
0.49 |
Binding ≤ 10μM
|
|
NOS3-3-E |
Nitric Oxide Synthase, Endothelial (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
180 |
0.47 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.73 |
8.38 |
-39.13 |
1 |
5 |
1 |
53 |
298.794 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.73 |
8.03 |
-35.47 |
1 |
5 |
1 |
53 |
298.794 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 55 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
1300 |
1.03 |
Binding ≤ 10μM
|
|
NOS2-8-E |
Nitric Oxide Synthase, Inducible (cluster #8 Of 9), Eukaryotic |
Eukaryotes |
940 |
1.05 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
1200 |
1.04 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.92 |
2.72 |
-27.36 |
3 |
2 |
1 |
40 |
109.152 |
0 |
↓
|
|
Mid
Mid (pH 6-8)
|
0.92 |
2.24 |
-4.62 |
2 |
2 |
0 |
39 |
108.144 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS2-4-E |
Nitric Oxide Synthase, Inducible (cluster #4 Of 9), Eukaryotic |
Eukaryotes |
2700 |
0.78 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
3300 |
0.77 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.76 |
4.01 |
-27.08 |
3 |
2 |
1 |
40 |
137.206 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.76 |
3.54 |
-4.53 |
2 |
2 |
0 |
39 |
136.198 |
1 |
↓
|
|
|
|
Analogs
-
39051638
-
Draw
Identity
99%
90%
80%
70%
Vendors
And 6 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
340 |
1.01 |
Binding ≤ 10μM
|
|
NOS2-8-E |
Nitric Oxide Synthase, Inducible (cluster #8 Of 9), Eukaryotic |
Eukaryotes |
810 |
0.95 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
600 |
0.97 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.35 |
3.17 |
-26.64 |
3 |
2 |
1 |
40 |
123.179 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.35 |
2.69 |
-4.67 |
2 |
2 |
0 |
39 |
122.171 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-1-E |
Nitric-oxide Synthase, Brain (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
120 |
0.39 |
Binding ≤ 10μM
|
|
NOS2-4-E |
Nitric Oxide Synthase, Inducible (cluster #4 Of 9), Eukaryotic |
Eukaryotes |
5000 |
0.30 |
Binding ≤ 10μM
|
|
NOS3-3-E |
Nitric Oxide Synthase, Endothelial (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
3500 |
0.31 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.31 |
9.82 |
-28.79 |
4 |
3 |
1 |
50 |
370.929 |
8 |
↓
|
|
Mid
Mid (pH 6-8)
|
4.18 |
9.36 |
-7.3 |
3 |
3 |
0 |
50 |
369.921 |
7 |
↓
|
|
Mid
Mid (pH 6-8)
|
4.31 |
11.22 |
-52.4 |
4 |
3 |
1 |
52 |
370.929 |
8 |
↓
|
|
|
|
Analogs
-
39048555
-
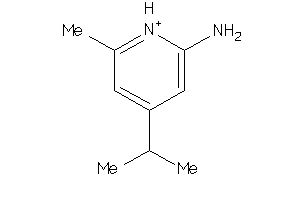
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS2-4-E |
Nitric Oxide Synthase, Inducible (cluster #4 Of 9), Eukaryotic |
Eukaryotes |
330 |
0.91 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
49 |
1.02 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.93 |
3.93 |
-24.72 |
3 |
2 |
1 |
40 |
137.206 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.93 |
3.56 |
-4.58 |
2 |
2 |
0 |
39 |
136.198 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 14 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.66 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
5500 |
0.49 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
320 |
0.61 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.63 |
1.88 |
-54.5 |
6 |
9 |
0 |
164 |
219.201 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.62 |
-49.85 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.58 |
-51.54 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
|
|
Analogs
-
39048555
-

Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
1200 |
0.75 |
Binding ≤ 10μM
|
|
NOS2-4-E |
Nitric Oxide Synthase, Inducible (cluster #4 Of 9), Eukaryotic |
Eukaryotes |
110 |
0.89 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
200 |
0.85 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.42 |
4.65 |
-24.01 |
3 |
2 |
1 |
40 |
151.233 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.42 |
3.98 |
-4.71 |
2 |
2 |
0 |
39 |
150.225 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
90 |
0.90 |
Binding ≤ 10μM
|
|
NOS2-4-E |
Nitric Oxide Synthase, Inducible (cluster #4 Of 9), Eukaryotic |
Eukaryotes |
15 |
1.00 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
50 |
0.93 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.44 |
4.71 |
-25.08 |
3 |
2 |
1 |
40 |
151.233 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.44 |
4.34 |
-4.4 |
2 |
2 |
0 |
39 |
150.225 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 2 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-3-E |
Nitric-oxide Synthase, Brain (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
3200 |
1.10 |
Binding ≤ 10μM
|
|
NOS2-2-E |
Nitric Oxide Synthase, Inducible (cluster #2 Of 9), Eukaryotic |
Eukaryotes |
2900 |
1.11 |
Binding ≤ 10μM
|
|
NOS3-5-E |
Nitric Oxide Synthase, Endothelial (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
7100 |
1.03 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.42 |
0.96 |
-26.9 |
3 |
2 |
1 |
38 |
117.197 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.42 |
0.92 |
-5.32 |
2 |
2 |
0 |
36 |
116.189 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
10000 |
0.54 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
340 |
0.70 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
540 |
0.67 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.39 |
1.4 |
-72.52 |
7 |
6 |
1 |
120 |
189.239 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
10000 |
0.54 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
340 |
0.70 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
540 |
0.67 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.39 |
-6.24 |
-77.63 |
7 |
6 |
1 |
119 |
189.239 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 23 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-3-E |
Nitric-oxide Synthase, Brain (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
29 |
1.76 |
Binding ≤ 10μM
|
|
NOS2-2-E |
Nitric Oxide Synthase, Inducible (cluster #2 Of 9), Eukaryotic |
Eukaryotes |
160 |
1.59 |
Binding ≤ 10μM
|
|
NOS3-5-E |
Nitric Oxide Synthase, Endothelial (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
1300 |
1.37 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.29 |
0.49 |
-28.44 |
4 |
2 |
1 |
52 |
105.186 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.29 |
0.6 |
-3.56 |
3 |
2 |
0 |
50 |
104.178 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
7600 |
0.48 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.63 |
2.42 |
-84.81 |
7 |
9 |
1 |
168 |
220.209 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
2.1 |
-57.3 |
6 |
9 |
0 |
167 |
219.201 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 76 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-3-E |
Nitric-oxide Synthase, Brain (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
160 |
1.90 |
Binding ≤ 10μM
|
|
NOS2-6-E |
Nitric Oxide Synthase, Inducible (cluster #6 Of 9), Eukaryotic |
Eukaryotes |
3000 |
1.55 |
Binding ≤ 10μM
|
|
NOS3-5-E |
Nitric Oxide Synthase, Endothelial (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
7000 |
1.44 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.09 |
-0.39 |
-28.46 |
4 |
2 |
1 |
52 |
91.159 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
-0.09 |
-0.26 |
-3.76 |
3 |
2 |
0 |
50 |
90.151 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 46 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-3-E |
Nitric-oxide Synthase, Brain (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
410 |
1.49 |
Binding ≤ 10μM
|
|
NOS2-2-E |
Nitric Oxide Synthase, Inducible (cluster #2 Of 9), Eukaryotic |
Eukaryotes |
900 |
1.41 |
Binding ≤ 10μM
|
|
NOS3-5-E |
Nitric Oxide Synthase, Endothelial (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
1100 |
1.39 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.02 |
0.66 |
-27.86 |
3 |
2 |
1 |
40 |
103.17 |
0 |
↓
|
|
Ref
Reference (pH 7)
|
0.15 |
0.57 |
-27.91 |
3 |
2 |
1 |
38 |
103.17 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.15 |
0.57 |
-7.03 |
2 |
2 |
0 |
36 |
102.162 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
140 |
1.07 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
160 |
1.06 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
6300 |
0.81 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.36 |
2.59 |
-29 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
140 |
1.07 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
160 |
1.06 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
6300 |
0.81 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.36 |
2.73 |
-29.02 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
140 |
1.07 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
160 |
1.06 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
6300 |
0.81 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.36 |
2.72 |
-28.98 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
140 |
1.07 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
160 |
1.06 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
6300 |
0.81 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.36 |
2.59 |
-29.03 |
3 |
2 |
1 |
40 |
127.211 |
1 |
↓
|
|
|
|
Analogs
-
2582068
-
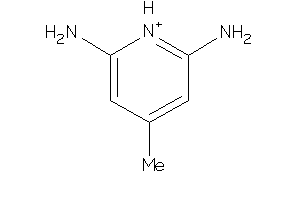
Draw
Identity
99%
90%
80%
70%
Vendors
And 67 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-5-E |
Nitric-oxide Synthase, Brain (cluster #5 Of 7), Eukaryotic |
Eukaryotes |
40 |
1.29 |
Binding ≤ 10μM
|
|
NOS2-8-E |
Nitric Oxide Synthase, Inducible (cluster #8 Of 9), Eukaryotic |
Eukaryotes |
170 |
1.18 |
Binding ≤ 10μM
|
|
NOS3-6-E |
Nitric Oxide Synthase, Endothelial (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
72 |
1.25 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.92 |
2.63 |
-27.41 |
3 |
2 |
1 |
40 |
109.152 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.92 |
2.15 |
-4.55 |
2 |
2 |
0 |
39 |
108.144 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-3-E |
Nitric-oxide Synthase, Brain (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
570 |
0.73 |
Binding ≤ 10μM
|
|
NOS3-3-E |
Nitric Oxide Synthase, Endothelial (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
2600 |
0.65 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.66 |
4.59 |
-31.72 |
3 |
2 |
1 |
38 |
201.702 |
3 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.54 |
4.62 |
-3.64 |
2 |
2 |
0 |
38 |
200.694 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-4-E |
Nitric-oxide Synthase, Brain (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
390 |
0.90 |
Binding ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 9), Eukaryotic |
Eukaryotes |
300 |
0.91 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
3500 |
0.76 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.95 |
3.6 |
-29.3 |
3 |
2 |
1 |
40 |
141.238 |
2 |
↓
|
|