|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-1-F |
Tyrosinase (cluster #1 Of 8), Fungal |
Fungi |
970 |
0.35 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.03 |
10.58 |
-37.6 |
1 |
4 |
1 |
35 |
340.472 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
4.03 |
8.07 |
-9.84 |
0 |
4 |
0 |
34 |
339.464 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-1-F |
Tyrosinase (cluster #1 Of 8), Fungal |
Fungi |
1340 |
0.34 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.97 |
5.57 |
-10.68 |
0 |
5 |
0 |
43 |
341.436 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.97 |
8.14 |
-39.99 |
1 |
5 |
1 |
45 |
342.444 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-8-F |
Tyrosinase (cluster #8 Of 8), Fungal |
Fungi |
2400 |
0.56 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.47 |
2.29 |
-10.13 |
4 |
5 |
0 |
80 |
208.246 |
1 |
↓
|
|
Mid
Mid (pH 6-8)
|
1.00 |
3.43 |
-40.93 |
3 |
5 |
-1 |
77 |
207.238 |
0 |
↓
|
|
|
|
Analogs
-
26513874
-
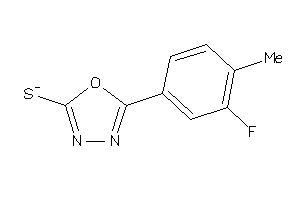
-
26514124
-
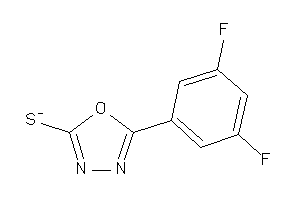
-
34964304
-
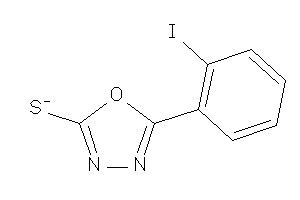
Draw
Identity
99%
90%
80%
70%
Vendors
And 46 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-1-F |
Tyrosinase (cluster #1 Of 8), Fungal |
Fungi |
6470 |
0.61 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.32 |
0.93 |
-42.39 |
0 |
3 |
-1 |
39 |
177.208 |
1 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.60 |
1.48 |
-7.53 |
1 |
3 |
0 |
42 |
178.216 |
1 |
↓
|
|
|
|
Analogs
-
26513878
-
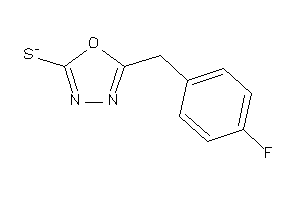
-
34964286
-
Draw
Identity
99%
90%
80%
70%
Vendors
And 21 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-1-F |
Tyrosinase (cluster #1 Of 8), Fungal |
Fungi |
3670 |
0.59 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.92 |
1.39 |
-39.88 |
0 |
3 |
-1 |
39 |
191.235 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
1.19 |
2.15 |
-10.85 |
1 |
3 |
0 |
42 |
192.243 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 23 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-3-F |
Tyrosinase (cluster #3 Of 8), Fungal |
Fungi |
400 |
1.00 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.59 |
2.97 |
-55.32 |
0 |
2 |
-1 |
40 |
121.115 |
0 |
↓
|
|
Mid
Mid (pH 6-8)
|
0.59 |
2.96 |
-8.86 |
1 |
2 |
0 |
37 |
122.123 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-4-F |
Tyrosinase (cluster #4 Of 8), Fungal |
Fungi |
290 |
0.83 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.13 |
0.77 |
-13.7 |
3 |
4 |
0 |
61 |
152.153 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.13 |
1.98 |
-54.46 |
2 |
4 |
-1 |
64 |
151.145 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-3-F |
Tyrosinase (cluster #3 Of 8), Fungal |
Fungi |
9530 |
0.59 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.58 |
4.86 |
-8.7 |
1 |
2 |
0 |
37 |
164.204 |
1 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.27 |
5.56 |
-47.62 |
0 |
2 |
-1 |
40 |
163.196 |
1 |
↓
|
|
Mid
Mid (pH 6-8)
|
1.58 |
5.66 |
-49.54 |
0 |
2 |
-1 |
40 |
163.196 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-3-F |
Tyrosinase (cluster #3 Of 8), Fungal |
Fungi |
70 |
0.83 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.10 |
0.79 |
-11.15 |
1 |
2 |
0 |
37 |
164.204 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-8-F |
Tyrosinase (cluster #8 Of 8), Fungal |
Fungi |
7700 |
0.42 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.52 |
2.29 |
-9.77 |
2 |
3 |
0 |
53 |
243.287 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
3.52 |
3.06 |
-45.66 |
1 |
3 |
-1 |
56 |
242.279 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-3-F |
Tyrosinase (cluster #3 Of 8), Fungal |
Fungi |
3800 |
0.84 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.78 |
1.81 |
-23.84 |
1 |
2 |
0 |
33 |
142.179 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-7-F |
Tyrosinase (cluster #7 Of 8), Fungal |
Fungi |
8800 |
0.29 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.74 |
-5.01 |
-15.88 |
4 |
5 |
0 |
85 |
322.364 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 11 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
TYRO-5-F |
Tyrosinase (cluster #5 Of 8), Fungal |
Fungi |
9000 |
0.54 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.02 |
4.52 |
-12.66 |
3 |
3 |
0 |
50 |
197.238 |
3 |
↓
|
|
Fatal error: Out of memory (allocated 12845056) (tried to allocate 163841 bytes) in /domains/zinc12/htdocs/tpl/render-item.tpl.php on line 1