|
|
Analogs
-
6116142
-
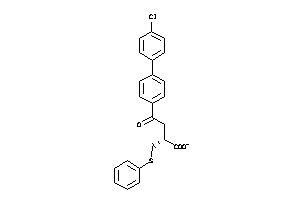
Draw
Identity
99%
90%
80%
70%
Vendors
And 5 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP1-1-E |
Matrix Metalloproteinase-1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
5000 |
0.27 |
Binding ≤ 10μM
|
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
1470 |
0.29 |
Binding ≤ 10μM
|
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
1570 |
0.29 |
Binding ≤ 10μM
|
|
MMP2-2-E |
72 KDa Type IV Collagenase (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
11 |
0.40 |
Binding ≤ 10μM
|
|
MMP2-2-E |
72 KDa Type IV Collagenase (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
91 |
0.35 |
Binding ≤ 10μM
|
|
MMP3-2-E |
Matrix Metalloproteinase 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
143 |
0.34 |
Binding ≤ 10μM
|
|
MMP3-2-E |
Matrix Metalloproteinase 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
7950 |
0.25 |
Binding ≤ 10μM
|
|
MMP8-4-E |
Matrix Metalloproteinase 8 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
303 |
0.33 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
301 |
0.33 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3110 |
0.28 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.88 |
1.92 |
-59.71 |
0 |
3 |
-1 |
57 |
409.914 |
8 |
↓
|
|
|
|
Analogs
-
538656
-
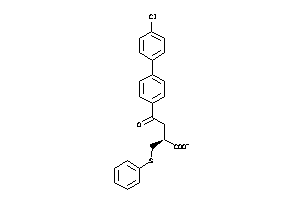
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
1570 |
0.29 |
Binding ≤ 10μM
|
|
MMP2-2-E |
72 KDa Type IV Collagenase (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
11 |
0.40 |
Binding ≤ 10μM
|
|
MMP2-2-E |
72 KDa Type IV Collagenase (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
91 |
0.35 |
Binding ≤ 10μM
|
|
MMP3-2-E |
Matrix Metalloproteinase 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
134 |
0.34 |
Binding ≤ 10μM
|
|
MMP3-2-E |
Matrix Metalloproteinase 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
7950 |
0.25 |
Binding ≤ 10μM
|
|
MMP8-4-E |
Matrix Metalloproteinase 8 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
303 |
0.33 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
301 |
0.33 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3110 |
0.28 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.88 |
2.11 |
-60.39 |
0 |
3 |
-1 |
57 |
409.914 |
8 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
73 |
0.24 |
Binding ≤ 10μM
|
|
MMP14-1-E |
Matrix Metalloproteinase 14 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
1900 |
0.20 |
Binding ≤ 10μM
|
|
MMP2-1-E |
72 KDa Type IV Collagenase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
1790 |
0.20 |
Binding ≤ 10μM
|
|
MMP3-1-E |
Matrix Metalloproteinase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
6 |
0.28 |
Binding ≤ 10μM
|
|
MMP9-2-E |
Matrix Metalloproteinase 9 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
840 |
0.21 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
6.37 |
-1.05 |
-17.51 |
4 |
7 |
0 |
107 |
557.735 |
13 |
↓
|
|
Hi
High (pH 8-9.5)
|
6.55 |
-0.48 |
-57.77 |
3 |
7 |
-1 |
113 |
556.727 |
13 |
↓
|
|
Hi
High (pH 8-9.5)
|
6.37 |
-0.48 |
-57.77 |
3 |
7 |
-1 |
110 |
556.727 |
13 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
2300 |
0.19 |
Binding ≤ 10μM
|
|
MMP3-1-E |
Matrix Metalloproteinase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
23 |
0.25 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
6.43 |
0.72 |
-50.23 |
2 |
7 |
-1 |
107 |
571.738 |
15 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP1-1-E |
Matrix Metalloproteinase-1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2200 |
0.33 |
Binding ≤ 10μM
|
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
11 |
0.46 |
Binding ≤ 10μM
|
|
MMP2-1-E |
72 KDa Type IV Collagenase (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2 |
0.51 |
Binding ≤ 10μM
|
|
MMP3-1-E |
Matrix Metalloproteinase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3 |
0.50 |
Binding ≤ 10μM
|
|
MMP7-1-E |
Matrix Metalloproteinase 7 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4500 |
0.31 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3900 |
0.32 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.15 |
7.61 |
-46.41 |
1 |
5 |
-1 |
86 |
346.428 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.35 |
Binding ≤ 10μM
|
|
MMP8-4-E |
Matrix Metalloproteinase 8 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
670 |
0.41 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
160 |
0.45 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.34 |
-1.38 |
-51.34 |
2 |
5 |
-1 |
92 |
292.355 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.35 |
Binding ≤ 10μM
|
|
MMP8-4-E |
Matrix Metalloproteinase 8 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
670 |
0.41 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
160 |
0.45 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.34 |
6.38 |
-45.59 |
2 |
5 |
-1 |
92 |
292.355 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
4700 |
0.31 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.37 |
6.33 |
-17.86 |
1 |
7 |
0 |
86 |
320.356 |
4 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.37 |
6.79 |
-47.51 |
2 |
7 |
1 |
87 |
321.364 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 8 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
3600 |
0.33 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.14 |
0.98 |
-40.34 |
1 |
6 |
-1 |
94 |
308.317 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
0.95 |
3.55 |
-9.43 |
2 |
6 |
0 |
88 |
309.325 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
0.58 |
1.04 |
-40.63 |
1 |
6 |
-1 |
94 |
308.317 |
4 |
↓
|
|
|
|
Analogs
-
4501578
-
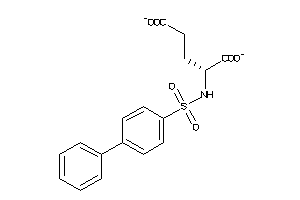
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
280 |
0.37 |
Binding ≤ 10μM
|
|
MMP8-3-E |
Matrix Metalloproteinase 8 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
1024 |
0.34 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.36 |
7.17 |
-107.22 |
1 |
7 |
-2 |
126 |
361.375 |
8 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.36 |
5.2 |
-49.29 |
2 |
7 |
-1 |
124 |
362.383 |
8 |
↓
|
|
|
|
Analogs
-
4501577
-
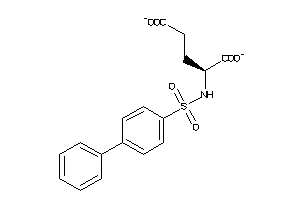
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATS4-1-E |
ADAMTS4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1100 |
0.33 |
Binding ≤ 10μM
|
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
280 |
0.37 |
Binding ≤ 10μM
|
|
MMP8-3-E |
Matrix Metalloproteinase 8 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
1024 |
0.34 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.36 |
7.1 |
-111.2 |
1 |
7 |
-2 |
126 |
361.375 |
8 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.36 |
5.13 |
-57.9 |
2 |
7 |
-1 |
124 |
362.383 |
8 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
9900 |
0.28 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.70 |
8.44 |
-17.08 |
0 |
6 |
0 |
81 |
359.403 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
2500 |
0.34 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.30 |
3.69 |
-7.42 |
2 |
6 |
0 |
88 |
315.373 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.16 |
1.16 |
-37.87 |
1 |
6 |
-1 |
94 |
314.365 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.60 |
1.2 |
-37.78 |
1 |
6 |
-1 |
94 |
314.365 |
4 |
↓
|
|
|
|
Analogs
-
11525611
-
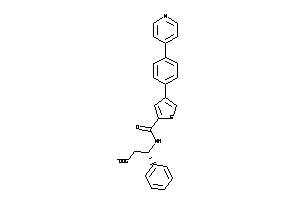
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MMP12-1-E |
Macrophage Metalloelastase (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
14 |
0.35 |
Binding ≤ 10μM
|
|
MMP13-4-E |
Matrix Metalloproteinase 13 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
270 |
0.30 |
Binding ≤ 10μM
|
|
MMP3-1-E |
Matrix Metalloproteinase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
390 |
0.29 |
Binding ≤ 10μM
|
|
MMP8-1-E |
Matrix Metalloproteinase 8 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1700 |
0.26 |
Binding ≤ 10μM
|
|
MMP9-1-E |
Matrix Metalloproteinase 9 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
980 |
0.27 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.46 |
-0.31 |
-56.05 |
1 |
5 |
-1 |
82 |
427.505 |
7 |
↓
|
|