|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
60 |
0.78 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
400 |
0.69 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.52 |
1.26 |
-72.12 |
6 |
5 |
1 |
105 |
206.291 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.65 |
0.87 |
-52.03 |
5 |
5 |
0 |
106 |
205.283 |
6 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.52 |
0.93 |
-52.68 |
5 |
5 |
0 |
104 |
205.283 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DDAH1-1-E |
N(G),N(G)-dimethylarginine Dimethylaminohydrolase 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
2000 |
0.57 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
100 |
0.70 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.07 |
1.88 |
-42.53 |
6 |
5 |
0 |
106 |
199.254 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.24 |
1.43 |
-53.82 |
5 |
5 |
0 |
106 |
199.254 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.07 |
0.61 |
-116.68 |
7 |
5 |
1 |
107 |
200.262 |
7 |
↓
|
|
|
|
Analogs
-
4556609
-
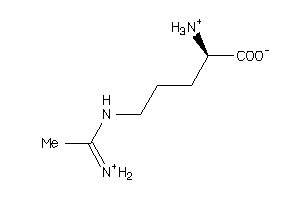
Draw
Identity
99%
90%
80%
70%
Vendors
And 5 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
9000 |
0.59 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
2200 |
0.66 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
4000 |
0.63 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
810 |
0.71 |
Binding ≤ 10μM |
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
1420 |
0.68 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.65 |
0.77 |
-74.95 |
6 |
5 |
1 |
105 |
174.224 |
6 |
↓
|
|
|
|
Analogs
-
19594743
-
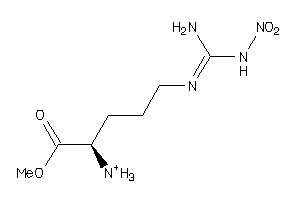
Draw
Identity
99%
90%
80%
70%
Vendors
And 41 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
690 |
0.54 |
Binding ≤ 10μM |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
1600 |
0.51 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
830 |
0.53 |
Binding ≤ 10μM |
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
8000 |
0.45 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
680 |
0.54 |
Binding ≤ 10μM |
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
2700 |
0.49 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
150 |
0.60 |
Functional ≤ 10μM
|
|
NOS3-1-E |
Nitric-oxide Synthase, Endothelial (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2700 |
0.49 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.56 |
1.67 |
-17.55 |
5 |
9 |
0 |
149 |
233.228 |
8 |
↓
|
|
Ref
Reference (pH 7)
|
-1.56 |
1.66 |
-19.79 |
5 |
9 |
0 |
149 |
233.228 |
8 |
↓
|
|
Mid
Mid (pH 6-8)
|
-1.56 |
1.79 |
-39.25 |
4 |
9 |
-1 |
151 |
232.22 |
8 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 8 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
4300 |
0.58 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
860 |
0.65 |
Binding ≤ 10μM |
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
4200 |
0.58 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
100 |
0.75 |
Functional ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Functional ≤ 10μM
|
|
NOS3-1-E |
Nitric-oxide Synthase, Endothelial (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Functional ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3000 |
0.59 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.26 |
1.39 |
-72.47 |
7 |
6 |
1 |
117 |
189.239 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
8.62 |
17.49 |
-47.16 |
0 |
0 |
-1 |
0 |
421.61 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
9600 |
0.78 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.81 |
3.32 |
-26.39 |
2 |
2 |
1 |
29 |
125.195 |
0 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-0.14 |
3.15 |
-109.21 |
4 |
2 |
2 |
32 |
126.203 |
0 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-0.14 |
3.38 |
-26.87 |
3 |
2 |
1 |
30 |
125.195 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
9600 |
0.78 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.81 |
3.36 |
-28.04 |
2 |
2 |
1 |
29 |
125.195 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 25 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.66 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
5500 |
0.49 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
7600 |
0.48 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
320 |
0.61 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3000 |
0.52 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.63 |
1.9 |
-54.25 |
6 |
9 |
0 |
164 |
219.201 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.57 |
-53.16 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.57 |
-48.12 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
100 |
0.58 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.58 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.18 |
-1.74 |
-126.95 |
10 |
9 |
2 |
164 |
249.319 |
10 |
↓
|
|
Mid
Mid (pH 6-8)
|
-3.18 |
-2.12 |
-112.39 |
9 |
9 |
1 |
166 |
248.311 |
10 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-3.18 |
-0.36 |
-249.23 |
11 |
9 |
3 |
168 |
250.327 |
10 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 14 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.66 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
5500 |
0.49 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
320 |
0.61 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.63 |
1.88 |
-54.5 |
6 |
9 |
0 |
164 |
219.201 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.62 |
-49.85 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.58 |
-51.54 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
10000 |
0.54 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
340 |
0.70 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
540 |
0.67 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.39 |
1.4 |
-72.52 |
7 |
6 |
1 |
120 |
189.239 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
10000 |
0.54 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
340 |
0.70 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
540 |
0.67 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.39 |
-6.24 |
-77.63 |
7 |
6 |
1 |
119 |
189.239 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
4300 |
0.58 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.72 |
1.31 |
-76.1 |
7 |
6 |
1 |
120 |
189.239 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
7600 |
0.48 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.63 |
2.42 |
-84.81 |
7 |
9 |
1 |
168 |
220.209 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
2.1 |
-57.3 |
6 |
9 |
0 |
167 |
219.201 |
7 |
↓
|
|
|
|
Analogs
-
4475245
-
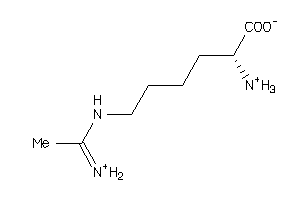
Draw
Identity
99%
90%
80%
70%
Vendors
And 6 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
6000 |
0.56 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
5000 |
0.57 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
7900 |
0.55 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
1600 |
0.62 |
Functional ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
130 |
0.74 |
Functional ≤ 10μM
|
|
NOS3-1-E |
Nitric-oxide Synthase, Endothelial (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2400 |
0.61 |
Functional ≤ 10μM
|
|
Z80120-5-O |
DLD-1 (Colon Adenocarcinoma Cells) (cluster #5 Of 5), Other |
Other |
2800 |
0.60 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.15 |
1.54 |
-74.91 |
6 |
5 |
1 |
105 |
188.251 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
60 |
0.84 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
3600 |
0.64 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.89 |
0.4 |
-54.94 |
6 |
5 |
0 |
106 |
191.256 |
6 |
↓
|
|