|
|
|
|
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DUS3-1-E |
Dual Specificity Protein Phosphatase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4200 |
0.22 |
Binding ≤ 10μM |
|
PTN7-1-E |
Protein-tyrosine Phosphatase LC-PTP (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1300 |
0.24 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.62 |
14.5 |
-48.94 |
0 |
8 |
-1 |
110 |
511.963 |
8 |
↓
|
|
|
|
Analogs
-
8425894
-
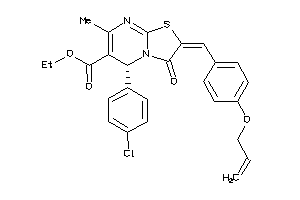
-
8425895
-
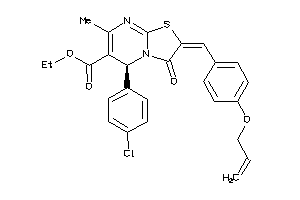
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DUS3-1-E |
Dual Specificity Protein Phosphatase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4200 |
0.22 |
Binding ≤ 10μM |
|
PTN7-1-E |
Protein-tyrosine Phosphatase LC-PTP (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1300 |
0.24 |
Binding ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.62 |
14.48 |
-48.78 |
0 |
8 |
-1 |
110 |
511.963 |
8 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DUS1-1-E |
Dual Specificity Protein Phosphatase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2800 |
0.19 |
Binding ≤ 10μM
|
|
DUS3-2-E |
Dual Specificity Protein Phosphatase 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
270 |
0.23 |
Binding ≤ 10μM
|
|
PTN1-1-E |
Protein-tyrosine Phosphatase 1B (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
2200 |
0.20 |
Binding ≤ 10μM
|
|
PTN7-1-E |
Protein-tyrosine Phosphatase LC-PTP (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
2400 |
0.20 |
Binding ≤ 10μM
|
|
PTPRC-3-E |
Leukocyte Common Antigen (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
1800 |
0.20 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.52 |
8.73 |
-53.3 |
0 |
10 |
-1 |
134 |
617.776 |
8 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DUS1-1-E |
Dual Specificity Protein Phosphatase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
460 |
0.22 |
Binding ≤ 10μM
|
|
DUS3-1-E |
Dual Specificity Protein Phosphatase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
18 |
0.27 |
Binding ≤ 10μM
|
|
MPIP1-2-E |
Dual Specificity Phosphatase Cdc25A (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
2400 |
0.20 |
Binding ≤ 10μM
|
|
PTN1-1-E |
Protein-tyrosine Phosphatase 1B (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
460 |
0.22 |
Binding ≤ 10μM
|
|
PTN7-1-E |
Protein-tyrosine Phosphatase LC-PTP (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
620 |
0.22 |
Binding ≤ 10μM
|
|
PTPRC-1-E |
Leukocyte Common Antigen (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
460 |
0.22 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.63 |
12.53 |
-51.54 |
0 |
8 |
-1 |
106 |
611.146 |
9 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DUS1-1-E |
Dual Specificity Protein Phosphatase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
780 |
0.22 |
Binding ≤ 10μM
|
|
DUS3-1-E |
Dual Specificity Protein Phosphatase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
78 |
0.26 |
Binding ≤ 10μM
|
|
MPIP1-2-E |
Dual Specificity Phosphatase Cdc25A (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
3400 |
0.20 |
Binding ≤ 10μM
|
|
PTN1-1-E |
Protein-tyrosine Phosphatase 1B (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1800 |
0.21 |
Binding ≤ 10μM
|
|
PTN7-1-E |
Protein-tyrosine Phosphatase LC-PTP (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1500 |
0.21 |
Binding ≤ 10μM
|
|
PTPRC-1-E |
Leukocyte Common Antigen (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
610 |
0.22 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.95 |
12.04 |
-51.46 |
0 |
8 |
-1 |
106 |
576.701 |
9 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DUS1-1-E |
Dual Specificity Protein Phosphatase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
470 |
0.22 |
Binding ≤ 10μM
|
|
DUS3-1-E |
Dual Specificity Protein Phosphatase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
71 |
0.25 |
Binding ≤ 10μM
|
|
MPIP1-2-E |
Dual Specificity Phosphatase Cdc25A (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
3400 |
0.19 |
Binding ≤ 10μM
|
|
PTN1-1-E |
Protein-tyrosine Phosphatase 1B (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
590 |
0.22 |
Binding ≤ 10μM
|
|
PTN7-1-E |
Protein-tyrosine Phosphatase LC-PTP (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1200 |
0.21 |
Binding ≤ 10μM
|
|
PTPRC-1-E |
Leukocyte Common Antigen (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
300 |
0.23 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.58 |
12.62 |
-50.63 |
0 |
8 |
-1 |
106 |
611.146 |
9 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DUS1-1-E |
Dual Specificity Protein Phosphatase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
520 |
0.22 |
Binding ≤ 10μM
|
|
DUS3-1-E |
Dual Specificity Protein Phosphatase 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
74 |
0.25 |
Binding ≤ 10μM
|
|
MPIP1-2-E |
Dual Specificity Phosphatase Cdc25A (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
2800 |
0.19 |
Binding ≤ 10μM
|
|
PTN1-1-E |
Protein-tyrosine Phosphatase 1B (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
420 |
0.22 |
Binding ≤ 10μM
|
|
PTN7-1-E |
Protein-tyrosine Phosphatase LC-PTP (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
870 |
0.21 |
Binding ≤ 10μM
|
|
PTPRC-1-E |
Leukocyte Common Antigen (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
500 |
0.22 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.07 |
12.21 |
-50.97 |
0 |
8 |
-1 |
106 |
594.691 |
9 |
↓
|
|