|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
9300 |
0.44 |
Binding ≤ 10μM
|
|
Z50643-2-O |
Hepatitis C Virus (cluster #2 Of 2), Other |
Other |
2600 |
0.49 |
Binding ≤ 10μM
|
|
Z50643-2-O |
Hepatitis C Virus (cluster #2 Of 5), Other |
Other |
9300 |
0.44 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.10 |
2.62 |
-54.11 |
2 |
6 |
-1 |
106 |
237.216 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.55 |
0.47 |
-115.71 |
1 |
6 |
-2 |
109 |
236.208 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
500 |
0.35 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.70 |
5.37 |
-45.68 |
0 |
5 |
-1 |
75 |
335.293 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
3.25 |
6.43 |
-21.15 |
1 |
5 |
0 |
72 |
336.301 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 2 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
1000 |
0.30 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.53 |
5.84 |
-53.88 |
1 |
7 |
-1 |
108 |
421.52 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
4000 |
0.29 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.27 |
-1.94 |
-63.78 |
1 |
8 |
-1 |
121 |
374.398 |
6 |
↓
|
|
|
|
Analogs
-
13590096
-
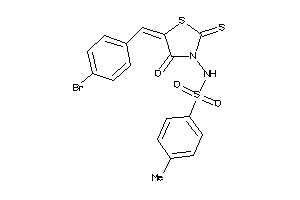
-
15986556
-
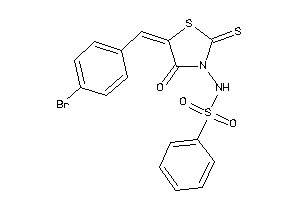
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
700 |
0.34 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.37 |
7.38 |
-16.46 |
1 |
5 |
0 |
68 |
455.38 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
3.37 |
8.3 |
-59.84 |
0 |
5 |
-1 |
70 |
454.372 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
2220 |
0.30 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.95 |
5.05 |
-131.52 |
0 |
7 |
-2 |
106 |
367.386 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.95 |
5.37 |
-56.09 |
1 |
7 |
-1 |
104 |
368.394 |
2 |
↓
|
|
|
|
Analogs
-
1213789
-
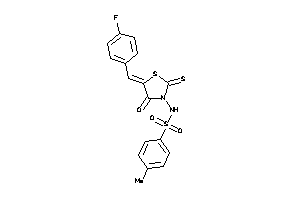
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
1000 |
0.34 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.72 |
6.83 |
-17.2 |
1 |
5 |
0 |
68 |
394.474 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.72 |
7.73 |
-60.75 |
0 |
5 |
-1 |
70 |
393.466 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
1550 |
0.30 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.45 |
5.81 |
-132.41 |
0 |
7 |
-2 |
106 |
381.413 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
5592 |
0.29 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.57 |
4.17 |
-129.57 |
0 |
7 |
-2 |
106 |
353.359 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
800 |
0.22 |
Binding ≤ 10μM
|
|
Q8JXU8-1-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #1 Of 1), Viral |
Viruses |
1100 |
0.21 |
Functional ≤ 10μM
|
|
Z100496-1-O |
Huh-5-2 (Huh-7 With Replicating HCV-RNA) (cluster #1 Of 1), Other |
Other |
1100 |
0.21 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
7.49 |
16.07 |
-63.78 |
0 |
6 |
-1 |
80 |
520.584 |
7 |
↓
|
|
Mid
Mid (pH 6-8)
|
7.49 |
16.37 |
-50.85 |
1 |
6 |
0 |
81 |
521.592 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
800 |
0.22 |
Binding ≤ 10μM
|
|
Q8JXU8-1-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #1 Of 1), Viral |
Viruses |
1100 |
0.21 |
Functional ≤ 10μM
|
|
Z100496-1-O |
Huh-5-2 (Huh-7 With Replicating HCV-RNA) (cluster #1 Of 1), Other |
Other |
1100 |
0.21 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
7.49 |
16.07 |
-63.69 |
0 |
6 |
-1 |
80 |
520.584 |
7 |
↓
|
|
Mid
Mid (pH 6-8)
|
7.49 |
16.37 |
-50.71 |
1 |
6 |
0 |
81 |
521.592 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
51 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.41 |
10.92 |
-56.49 |
0 |
5 |
-1 |
71 |
323.372 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
3.41 |
11.31 |
-26.06 |
1 |
5 |
0 |
72 |
324.38 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
80 |
0.40 |
Binding ≤ 10μM
|
|
Z50643-2-O |
Hepatitis C Virus (cluster #2 Of 5), Other |
Other |
2200 |
0.32 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.46 |
11.79 |
-57.59 |
0 |
4 |
-1 |
58 |
333.411 |
3 |
↓
|
|
|
|
Analogs
-
4526080
-
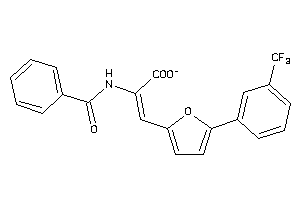
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
8700 |
0.24 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.70 |
11.54 |
-67.07 |
1 |
5 |
-1 |
82 |
400.332 |
6 |
↓
|
|
|
|
Analogs
-
1300993
-
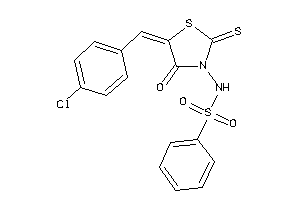
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
1500 |
0.33 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.24 |
8.19 |
-60.17 |
0 |
5 |
-1 |
70 |
409.921 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
4800 |
0.44 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.65 |
3.24 |
-37.12 |
0 |
4 |
-1 |
60 |
295.167 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.65 |
2.39 |
-6.59 |
1 |
4 |
0 |
58 |
296.175 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
80 |
0.34 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.52 |
7.37 |
-132.99 |
0 |
7 |
-2 |
106 |
409.467 |
5 |
↓
|
|
|
|
|
|
|
Analogs
-
15986554
-
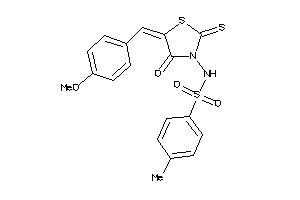
-
1300997
-
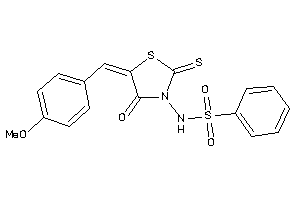
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
5000 |
0.29 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.62 |
5.85 |
-15.29 |
1 |
6 |
0 |
77 |
406.51 |
5 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.62 |
6.74 |
-62.41 |
0 |
6 |
-1 |
79 |
405.502 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
108 |
0.35 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.69 |
6.45 |
-132.03 |
0 |
7 |
-2 |
106 |
395.44 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.69 |
6.42 |
-57.22 |
1 |
7 |
-1 |
104 |
396.448 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
2200 |
0.20 |
Binding ≤ 10μM
|
|
Q8JXU8-1-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #1 Of 1), Viral |
Viruses |
2600 |
0.20 |
Functional ≤ 10μM
|
|
Z100496-1-O |
Huh-5-2 (Huh-7 With Replicating HCV-RNA) (cluster #1 Of 1), Other |
Other |
2600 |
0.20 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
6.26 |
14.29 |
-65.52 |
0 |
7 |
-1 |
93 |
521.572 |
7 |
↓
|
|
Mid
Mid (pH 6-8)
|
6.26 |
14.59 |
-53.84 |
1 |
7 |
0 |
94 |
522.58 |
7 |
↓
|
|
Lo
Low (pH 4.5-6)
|
6.26 |
15.03 |
-96.71 |
2 |
7 |
1 |
95 |
523.588 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
78 |
0.45 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.97 |
9.88 |
-52.06 |
0 |
3 |
-1 |
53 |
311.382 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
3400 |
0.33 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.19 |
10.13 |
-56.36 |
0 |
4 |
-1 |
58 |
325.413 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
1600 |
0.35 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.35 |
9.17 |
-58.38 |
0 |
5 |
-1 |
71 |
309.345 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Q8JXU8-2-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #2 Of 2), Viral |
Viruses |
200 |
0.33 |
Binding ≤ 10μM
|
|
Q8JXU8-1-V |
Hepatitis C Virus NS5B RNA-dependent RNA Polymerase (cluster #1 Of 1), Viral |
Viruses |
444 |
0.32 |
Functional ≤ 10μM
|
|
Z80169-1-O |
Huh-7 (Hepatocellular Carcinoma) (cluster #1 Of 1), Other |
Other |
444 |
0.32 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.01 |
6.59 |
-132.83 |
0 |
7 |
-2 |
106 |
395.44 |
4 |
↓
|
|