|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
98 |
0.75 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.79 |
2.2 |
-45.61 |
3 |
3 |
1 |
50 |
199.661 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
13 |
0.79 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.16 |
4.08 |
-40.96 |
2 |
3 |
1 |
39 |
213.688 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACH10-1-E |
Neuronal Acetylcholine Receptor Protein Alpha-10 Subunit (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA-2-E |
Acetylcholine Receptor Protein Alpha Chain (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA2-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-2 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
17 |
0.91 |
Binding ≤ 10μM
|
|
ACHA3-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-3 Subunit (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
2 |
1.01 |
Binding ≤ 10μM
|
|
ACHA5-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-5 Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA6-2-E |
Neuronal Acetylcholine Receptor Subunit Alpha-6 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA7-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-7 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA9-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-9 Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHB-4-E |
Acetylcholine Receptor Protein Beta Chain (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHB2-2-E |
Neuronal Acetylcholine Receptor Protein Beta-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
17 |
0.91 |
Binding ≤ 10μM
|
|
ACHB3-2-E |
Neuronal Acetylcholine Receptor Subunit Beta-3 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHB4-3-E |
Neuronal Acetylcholine Receptor Subunit Beta-4 (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
17 |
0.91 |
Binding ≤ 10μM
|
|
ACHD-3-E |
Acetylcholine Receptor Protein Delta Chain (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
700 |
0.72 |
Binding ≤ 10μM
|
|
ACHA3-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-3 Subunit (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
700 |
0.72 |
Functional ≤ 10μM
|
|
ACHA7-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-7 Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
8900 |
0.59 |
Functional ≤ 10μM
|
|
Z104281-2-O |
Neuronal Acetylcholine Receptor; Alpha3/beta2 (cluster #2 Of 2), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
700 |
0.72 |
Binding ≤ 10μM
|
|
Z104286-2-O |
Neuronal Acetylcholine Receptor; Alpha2/beta2 (cluster #2 Of 2), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104287-3-O |
Neuronal Acetylcholine Receptor; Alpha3/beta4 (cluster #3 Of 3), Other |
Other |
72 |
0.83 |
Binding ≤ 10μM
|
|
Z104287-3-O |
Neuronal Acetylcholine Receptor; Alpha3/beta4 (cluster #3 Of 3), Other |
Other |
74 |
0.83 |
Binding ≤ 10μM
|
|
Z104289-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta4 (cluster #2 Of 2), Other |
Other |
7 |
0.95 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
2 |
1.01 |
Binding ≤ 10μM
|
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
3 |
0.99 |
Functional ≤ 10μM
|
|
Z80175-1-O |
IMR-32 (Neuroblastoma Cells) (cluster #1 Of 2), Other |
Other |
700 |
0.72 |
Functional ≤ 10μM
|
|
Z81074-1-O |
K177cell Line (cluster #1 Of 1), Other |
Other |
700 |
0.72 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.83 |
-2.56 |
-40.53 |
2 |
3 |
1 |
38 |
165.216 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.83 |
-2.45 |
-88.84 |
3 |
3 |
2 |
39 |
166.224 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
2 |
0.68 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
2 |
0.68 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.63 |
1.67 |
-45.32 |
2 |
5 |
1 |
51 |
252.338 |
5 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.63 |
1.96 |
-89.09 |
3 |
5 |
2 |
52 |
253.346 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
2 |
0.68 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.86 |
3.88 |
-43.87 |
2 |
4 |
1 |
42 |
250.366 |
4 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.86 |
4.17 |
-87.91 |
3 |
4 |
2 |
43 |
251.374 |
4 |
↓
|
|
|
|
Analogs
-
13704072
-
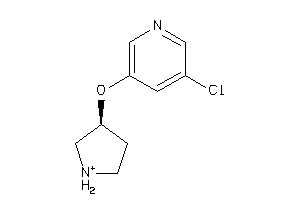
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
220 |
0.72 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.65 |
2.98 |
-44.64 |
2 |
3 |
1 |
39 |
199.661 |
2 |
↓
|
|
|
|
Analogs
-
13704068
-
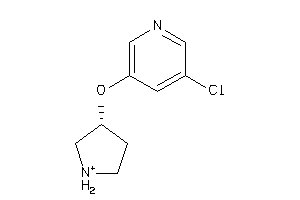
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
220 |
0.72 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.65 |
2.98 |
-43.56 |
2 |
3 |
1 |
39 |
199.661 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA2-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-2 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
320 |
0.65 |
Binding ≤ 10μM
|
|
ACHA3-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-3 Subunit (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
8 |
0.81 |
Binding ≤ 10μM
|
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
220 |
0.67 |
Binding ≤ 10μM
|
|
ACHA7-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-7 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
1530 |
0.58 |
Binding ≤ 10μM
|
|
ACHB2-2-E |
Neuronal Acetylcholine Receptor Protein Beta-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
8 |
0.81 |
Binding ≤ 10μM
|
|
ACHB4-3-E |
Neuronal Acetylcholine Receptor Subunit Beta-4 (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
320 |
0.65 |
Binding ≤ 10μM
|
|
ACHD-3-E |
Acetylcholine Receptor Protein Delta Chain (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
750 |
0.61 |
Binding ≤ 10μM
|
|
Z104287-3-O |
Neuronal Acetylcholine Receptor; Alpha3/beta4 (cluster #3 Of 3), Other |
Other |
1400 |
0.59 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
2 |
0.87 |
Binding ≤ 10μM
|
|
Z104283-1-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #1 Of 2), Other |
Other |
750 |
0.61 |
Functional ≤ 10μM |
|
Z81074-1-O |
K177cell Line (cluster #1 Of 1), Other |
Other |
750 |
0.61 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.34 |
0.08 |
-37.62 |
1 |
3 |
1 |
26 |
193.27 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
1300 |
0.55 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.04 |
3.79 |
-37.59 |
1 |
4 |
1 |
39 |
228.703 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.04 |
1.24 |
-5.87 |
0 |
4 |
0 |
38 |
227.695 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.41 |
2.38 |
-41.47 |
2 |
4 |
1 |
48 |
209.269 |
5 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.41 |
2.8 |
-94.02 |
3 |
4 |
2 |
49 |
210.277 |
5 |
↓
|
|
|
|
Analogs
-
13704014
-
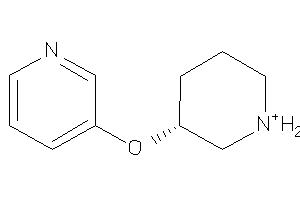
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
191 |
0.72 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
204 |
0.72 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
190 |
0.72 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.75 |
3.1 |
-46.36 |
2 |
3 |
1 |
39 |
179.243 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.75 |
3.38 |
-91.65 |
3 |
3 |
2 |
40 |
180.251 |
2 |
↓
|
|
|
|
Analogs
-
13704010
-
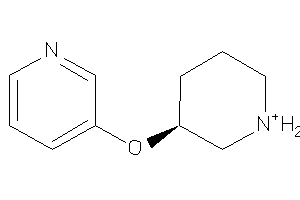
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
191 |
0.72 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
204 |
0.72 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
190 |
0.72 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.75 |
3.1 |
-46.38 |
2 |
3 |
1 |
39 |
179.243 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.75 |
3.38 |
-91.68 |
3 |
3 |
2 |
40 |
180.251 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 5 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHD-3-E |
Acetylcholine Receptor Protein Delta Chain (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.06 |
3.43 |
-40.62 |
2 |
3 |
1 |
39 |
291.112 |
3 |
↓
|
|
|
|
Analogs
-
18379
-
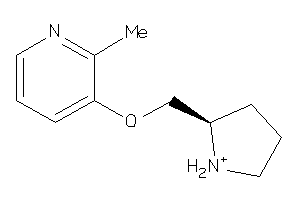
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHD-3-E |
Acetylcholine Receptor Protein Delta Chain (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
17 |
0.78 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
19 |
0.77 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
39 |
0.74 |
Binding ≤ 10μM
|
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
1100 |
0.60 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.32 |
4 |
-39.9 |
2 |
3 |
1 |
39 |
193.27 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.32 |
4.26 |
-84.98 |
3 |
3 |
2 |
40 |
194.278 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA2-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-2 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
4 |
0.56 |
Binding ≤ 10μM
|
|
ACHA3-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-3 Subunit (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
14 |
0.52 |
Binding ≤ 10μM
|
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
2000 |
0.38 |
Binding ≤ 10μM
|
|
ACHB2-2-E |
Neuronal Acetylcholine Receptor Protein Beta-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
4 |
0.56 |
Binding ≤ 10μM
|
|
ACHB4-3-E |
Neuronal Acetylcholine Receptor Subunit Beta-4 (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
3700 |
0.36 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
1 |
0.60 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.87 |
10.13 |
-37.67 |
1 |
3 |
1 |
27 |
291.39 |
6 |
↓
|
|
Lo
Low (pH 4.5-6)
|
2.87 |
10.4 |
-90.88 |
2 |
3 |
2 |
28 |
292.398 |
6 |
↓
|
|
|
|
Analogs
-
13704006
-
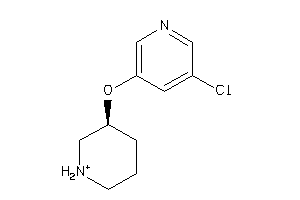
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
240 |
0.66 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
240 |
0.66 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.57 |
3.58 |
-44.15 |
2 |
3 |
1 |
39 |
213.688 |
2 |
↓
|
|
|
|
Analogs
-
13704002
-
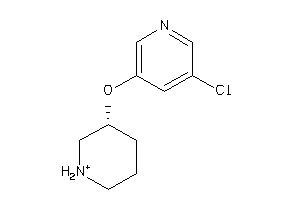
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
240 |
0.66 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
240 |
0.66 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.57 |
3.58 |
-44.12 |
2 |
3 |
1 |
39 |
213.688 |
2 |
↓
|
|
|
|
Analogs
-
4638275
-
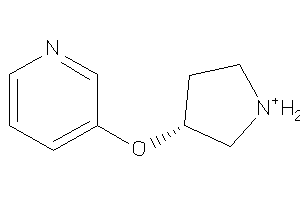
Draw
Identity
99%
90%
80%
70%
Vendors
And 16 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
100 |
0.82 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
209 |
0.78 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
449 |
0.74 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
100 |
0.82 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.83 |
2.47 |
-45.43 |
2 |
3 |
1 |
39 |
165.216 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.83 |
2.75 |
-89.25 |
3 |
3 |
2 |
40 |
166.224 |
2 |
↓
|
|
|
|
Analogs
-
3932135
-
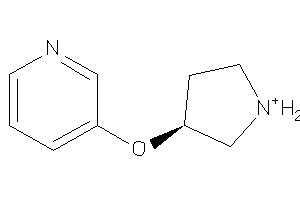
Draw
Identity
99%
90%
80%
70%
Vendors
And 16 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
100 |
0.82 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
45 |
0.86 |
Binding ≤ 10μM
|
|
Z104283-2-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #2 Of 4), Other |
Other |
209 |
0.78 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
100 |
0.82 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.83 |
2.44 |
-46.27 |
2 |
3 |
1 |
39 |
165.216 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.83 |
2.72 |
-90.43 |
3 |
3 |
2 |
40 |
166.224 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
500 |
0.59 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.84 |
1.73 |
-45.21 |
2 |
4 |
1 |
42 |
208.285 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.84 |
2.01 |
-91.88 |
3 |
4 |
2 |
43 |
209.293 |
3 |
↓
|
|
|
|
Analogs
-
26717002
-
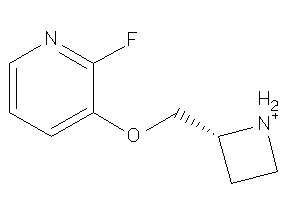
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACH10-1-E |
Neuronal Acetylcholine Receptor Protein Alpha-10 Subunit (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA-2-E |
Acetylcholine Receptor Protein Alpha Chain (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA2-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-2 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA2-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-2 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
181 |
0.73 |
Binding ≤ 10μM
|
|
ACHA3-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-3 Subunit (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA3-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-3 Subunit (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
3680 |
0.59 |
Binding ≤ 10μM
|
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
188 |
0.72 |
Binding ≤ 10μM
|
|
ACHA5-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-5 Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA6-2-E |
Neuronal Acetylcholine Receptor Subunit Alpha-6 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA7-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-7 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHA9-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-9 Subunit (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
ACHD-3-E |
Acetylcholine Receptor Protein Delta Chain (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.11 |
-1.94 |
-44.73 |
2 |
3 |
1 |
38 |
183.206 |
3 |
↓
|
|
|
|
Analogs
-
6525315
-
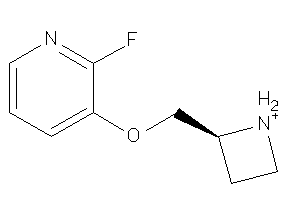
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHD-3-E |
Acetylcholine Receptor Protein Delta Chain (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.12 |
2.18 |
-44.24 |
2 |
3 |
1 |
39 |
183.206 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
0 |
0.00 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.12 |
-2.88 |
-43.33 |
2 |
4 |
1 |
51 |
166.204 |
3 |
↓
|
|
|
|
Analogs
-
6562
-
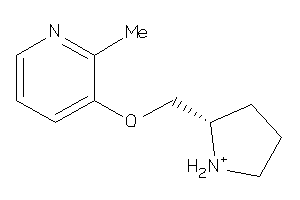
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHD-3-E |
Acetylcholine Receptor Protein Delta Chain (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
39 |
0.74 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
39 |
0.74 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.32 |
3.95 |
-39.66 |
2 |
3 |
1 |
39 |
193.27 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.32 |
4.22 |
-85.01 |
3 |
3 |
2 |
40 |
194.278 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACHA2-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-2 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
750 |
0.54 |
Binding ≤ 10μM
|
|
ACHA3-2-E |
Neuronal Acetylcholine Receptor Protein Alpha-3 Subunit (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
8 |
0.71 |
Binding ≤ 10μM
|
|
ACHA4-3-E |
Neuronal Acetylcholine Receptor Protein Alpha-4 Subunit (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
710 |
0.54 |
Binding ≤ 10μM
|
|
ACHB2-2-E |
Neuronal Acetylcholine Receptor Protein Beta-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
8 |
0.71 |
Binding ≤ 10μM
|
|
ACHB4-3-E |
Neuronal Acetylcholine Receptor Subunit Beta-4 (cluster #3 Of 7), Eukaryotic |
Eukaryotes |
750 |
0.54 |
Binding ≤ 10μM
|
|
Z104287-3-O |
Neuronal Acetylcholine Receptor; Alpha3/beta4 (cluster #3 Of 3), Other |
Other |
6800 |
0.45 |
Binding ≤ 10μM
|
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
2 |
0.76 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.09 |
6.7 |
-35.99 |
1 |
3 |
1 |
27 |
217.292 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.09 |
6.97 |
-87.3 |
2 |
3 |
2 |
28 |
218.3 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
500 |
0.63 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.47 |
0.8 |
-45.46 |
2 |
4 |
1 |
42 |
194.258 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.47 |
1.09 |
-91.35 |
3 |
4 |
2 |
43 |
195.266 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
1010 |
0.60 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.52 |
4.51 |
-36.8 |
2 |
3 |
1 |
39 |
193.27 |
5 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.52 |
4.93 |
-85.92 |
3 |
3 |
2 |
40 |
194.278 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
273 |
0.61 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.34 |
6.26 |
-34.26 |
1 |
3 |
1 |
27 |
272.166 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.34 |
3.93 |
-3.86 |
0 |
3 |
0 |
25 |
271.158 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
304 |
0.61 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.41 |
6.08 |
-36.62 |
1 |
3 |
1 |
27 |
227.715 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.41 |
3.75 |
-4.79 |
0 |
3 |
0 |
25 |
226.707 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
394 |
0.60 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.54 |
6.16 |
-36.54 |
1 |
3 |
1 |
27 |
272.166 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.54 |
3.84 |
-4.74 |
0 |
3 |
0 |
25 |
271.158 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
3 |
0.92 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.14 |
3.62 |
-36.62 |
2 |
3 |
1 |
39 |
179.243 |
4 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.14 |
3.89 |
-84.72 |
3 |
3 |
2 |
40 |
180.251 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
1 |
0.90 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.10 |
4.26 |
-38.58 |
2 |
3 |
1 |
39 |
258.139 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
266 |
0.71 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.82 |
2.41 |
-42.71 |
3 |
3 |
1 |
50 |
179.243 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.82 |
2.79 |
-89.19 |
4 |
3 |
2 |
51 |
180.251 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
40 |
0.74 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.20 |
4.27 |
-38.24 |
2 |
3 |
1 |
39 |
193.27 |
4 |
↓
|
|
Lo
Low (pH 4.5-6)
|
1.20 |
4.66 |
-84.57 |
3 |
3 |
2 |
40 |
194.278 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104290-4-O |
Neuronal Acetylcholine Receptor; Alpha4/beta2 (cluster #4 Of 4), Other |
Other |
39 |
0.74 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.30 |
4.17 |
-40.96 |
2 |
3 |
1 |
39 |
258.139 |
4 |
↓
|
|