|
|
Analogs
-
36473263
-
Draw
Identity
99%
90%
80%
70%
Vendors
And 29 More
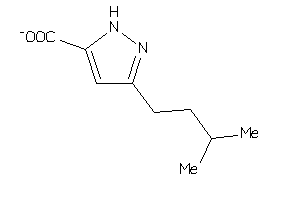
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.43 |
2.68 |
-42.89 |
1 |
4 |
-1 |
69 |
153.161 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 7 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104302-3-O |
Glutamate NMDA Receptor (cluster #3 Of 7), Other |
Other |
10000 |
0.64 |
Binding ≤ 10μM
|
|
Z50597-8-O |
Rattus Norvegicus (cluster #8 Of 12), Other |
Other |
9600 |
0.64 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.16 |
-2.03 |
-38.98 |
3 |
6 |
-1 |
112 |
157.105 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.70 |
-3.63 |
-100.85 |
2 |
6 |
-2 |
115 |
156.097 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-2.70 |
-4.07 |
-38.9 |
4 |
6 |
0 |
114 |
158.113 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2430 |
0.71 |
Binding ≤ 10μM |
|
GRIK2-3-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
3230 |
0.70 |
Binding ≤ 10μM |
|
GRIK3-3-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
1740 |
0.73 |
Binding ≤ 10μM |
|
GRM1-2-E |
Metabotropic Glutamate Receptor 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.66 |
Binding ≤ 10μM
|
|
GRM3-1-E |
Metabotropic Glutamate Receptor 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
10000 |
0.64 |
Binding ≤ 10μM
|
|
GRM5-2-E |
Metabotropic Glutamate Receptor 5 (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
8000 |
0.65 |
Binding ≤ 10μM
|
|
GRM1-3-E |
Metabotropic Glutamate Receptor 1 (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.66 |
Functional ≤ 10μM
|
|
GRM5-4-E |
Metabotropic Glutamate Receptor 5 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
6000 |
0.66 |
Functional ≤ 10μM
|
|
Z104302-3-O |
Glutamate NMDA Receptor (cluster #3 Of 7), Other |
Other |
10000 |
0.64 |
Binding ≤ 10μM
|
|
Z50597-8-O |
Rattus Norvegicus (cluster #8 Of 12), Other |
Other |
9600 |
0.64 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.16 |
-1.99 |
-41.18 |
3 |
6 |
-1 |
112 |
157.105 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.70 |
-3.67 |
-101.23 |
2 |
6 |
-2 |
115 |
156.097 |
2 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-2.70 |
-4.09 |
-41.28 |
4 |
6 |
0 |
114 |
158.113 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 11 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
5.09 |
2.22 |
-55.88 |
0 |
3 |
-1 |
53 |
374.366 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.64 |
3.01 |
-97.1 |
9 |
10 |
2 |
162 |
509.586 |
6 |
↓
|
|
|
|
Analogs
-
36473263
-

Draw
Identity
99%
90%
80%
70%
Vendors
And 23 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.99 |
3.46 |
-42.89 |
1 |
4 |
-1 |
69 |
167.188 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 94 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
HCAR2-2-E |
HM74 Nicotinic Acid GPCR (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
270 |
1.02 |
Binding ≤ 10μM
|
|
HCAR2-2-E |
HM74 Nicotinic Acid GPCR (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
29 |
1.17 |
Functional ≤ 10μM |
|
Z50588-6-O |
Canis Familiaris (cluster #6 Of 7), Other |
Other |
2700 |
0.87 |
Functional ≤ 10μM
|
|
Z50597-8-O |
Rattus Norvegicus (cluster #8 Of 12), Other |
Other |
960 |
0.94 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.27 |
3.29 |
-45.83 |
0 |
3 |
-1 |
53 |
122.103 |
1 |
↓
|
|
Lo
Low (pH 4.5-6)
|
0.27 |
3.74 |
-46.83 |
1 |
3 |
0 |
54 |
123.111 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 59 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.79 |
2.47 |
-47.2 |
1 |
4 |
-1 |
69 |
151.145 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIA4-2-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2200 |
0.40 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
Z104302-4-O |
Glutamate NMDA Receptor (cluster #4 Of 7), Other |
Other |
7400 |
0.36 |
Binding ≤ 10μM |
|
Z50574-1-O |
Xenopus Sp. (cluster #1 Of 2), Other |
Other |
260 |
0.46 |
Functional ≤ 10μM
|
|
Z50597-8-O |
Rattus Norvegicus (cluster #8 Of 12), Other |
Other |
810 |
0.43 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.26 |
2.06 |
-40.89 |
1 |
9 |
-1 |
132 |
272.2 |
2 |
↓
|
|
Ref
Reference (pH 7)
|
0.22 |
5.15 |
-137.07 |
2 |
9 |
2 |
126 |
273.208 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.22 |
4.1 |
-49.04 |
1 |
9 |
1 |
125 |
272.2 |
2 |
↓
|
|