The SWEETLEAD database has been created to provide an exhaustive and highly curated resource for chemical structures of the world's approved medicines, illegal drugs, and isolates from traditional medicinal herbs. This database has been built using a consensus generating scheme pulling data from several public chemical databases (such as PubChem, ChemSpider, PharmGKB, etc.), as detailed in the publication. The paper is: Novick, P.A., Ortiz, O.F., Poelman, J., Abdulhay, A.Y., Pande, V.S. (2013) SWEETLEAD: An in silico database of approved drugs, regulated chemicals, and herbal isolates for computer-aided drug discovery. PLoS ONE (2013) We are grateful to the authors for creating and curating this database and thank them for allowing us to incorporate its structures in ZINC.
We assess the chemical diversity of a subset by clustering the molecules. First, we sort ligands by increasing molecular weight. Then, we use the SUBSET 1.0 algorithm ( Voigt JH, Bienfait B, Wang S, Nicklaus MC. JCICS, 2001, 41, 702-12) to progressively select compounds that differ from those previously selected by at least the Tanimoto cutoff, using ChemAxon default fingerprints. The resulting representatives have two interesting properties:
| Tanimoto Cutoff Level | 60% | 70% | 80% | 90% | 100% |
|---|---|---|---|---|---|
| Number of Representatives | 938 | 1,420 | 1,924 | 2,530 | 4,770 |
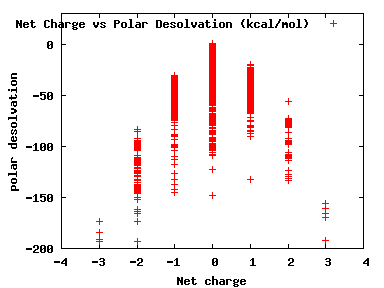
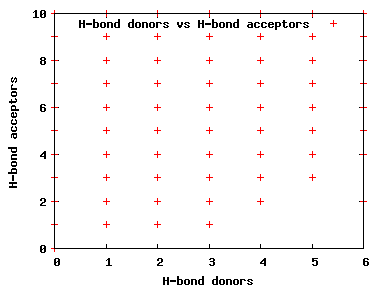
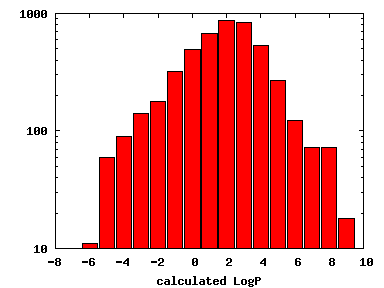
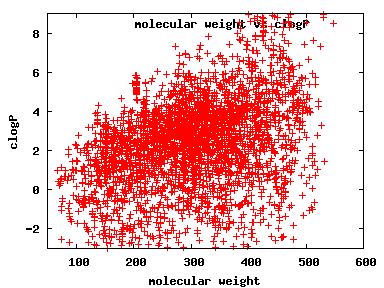
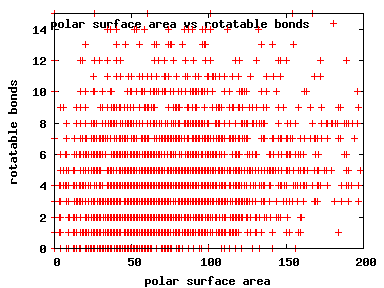
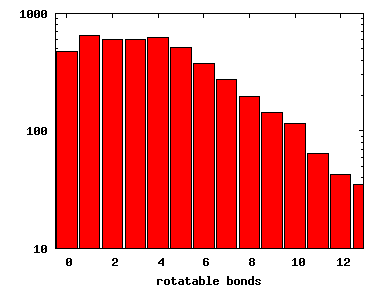
We compute the physical properties of each molecule in the subset, and graph them below.
Download Calculated Physical Properties
| Format | Reference(pH 7) | Mid(pH 6-8) | High(pH 8-9.5) | Low(pH 4.5-6) | Download Unix |
Download Windows |
|---|---|---|---|---|---|---|
| SMILES | All | All | All | All | ||
| MOL2 | All | All | All | All | Single Usual Metals All | Single Usual Metals All |
| SDF | All | All | All | All | Single Usual Metals All | Single Usual Metals All |
| Flexibase | Not Available | Not Available | Not Available | Not Available |