|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
9000 |
0.44 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
9000 |
0.44 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.09 |
5.74 |
-11.54 |
4 |
4 |
0 |
77 |
252.73 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.23 |
6.22 |
-37.51 |
5 |
4 |
1 |
77 |
253.738 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
9000 |
0.37 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
9000 |
0.37 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.83 |
5.43 |
-11.88 |
5 |
5 |
0 |
93 |
271.349 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
1.97 |
7.14 |
-37.1 |
6 |
5 |
1 |
92 |
272.357 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1600 |
0.43 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1600 |
0.43 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.04 |
5.6 |
-14.97 |
5 |
5 |
0 |
93 |
271.349 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.17 |
5.97 |
-38.66 |
6 |
5 |
1 |
92 |
272.357 |
3 |
↓
|
|
|
|
Analogs
-
4676424
-
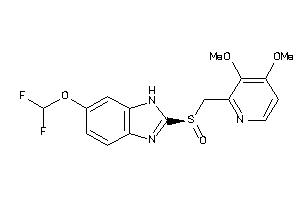
Draw
Identity
99%
90%
80%
70%
Vendors
And 41 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1000 |
0.32 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1000 |
0.32 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.95 |
2.48 |
-24.9 |
1 |
7 |
0 |
86 |
383.376 |
7 |
↓
|
|
Ref
Reference (pH 7)
|
1.95 |
2.49 |
-24.17 |
1 |
7 |
0 |
86 |
383.376 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.95 |
2.02 |
-54.17 |
0 |
7 |
-1 |
85 |
382.368 |
7 |
↓
|
|
|
|
Analogs
-
4099200
-
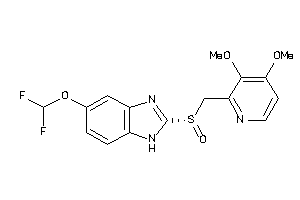
Draw
Identity
99%
90%
80%
70%
Vendors
And 37 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1000 |
0.32 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1000 |
0.32 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.95 |
2.48 |
-16.82 |
1 |
7 |
0 |
86 |
383.376 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.95 |
2.01 |
-55.16 |
0 |
7 |
-1 |
85 |
382.368 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.95 |
2.01 |
-55.75 |
0 |
7 |
-1 |
85 |
382.368 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 79 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
501 |
0.37 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
501 |
0.37 |
Binding ≤ 10μM
|
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
370 |
0.38 |
Functional ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
370 |
0.38 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.41 |
4.06 |
-43.78 |
2 |
6 |
1 |
78 |
346.432 |
5 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.41 |
3.62 |
-13.05 |
1 |
6 |
0 |
77 |
345.424 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 87 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
501 |
0.37 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
501 |
0.37 |
Binding ≤ 10μM
|
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
370 |
0.38 |
Functional ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
370 |
0.38 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.41 |
3.99 |
-43.94 |
2 |
6 |
1 |
78 |
346.432 |
5 |
↓
|
|
Hi
High (pH 8-9.5)
|
2.41 |
3.56 |
-24.45 |
1 |
6 |
0 |
77 |
345.424 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.41 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.41 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.40 |
5.51 |
-10.88 |
1 |
4 |
0 |
49 |
299.399 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.41 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
700 |
0.41 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.40 |
5.72 |
-11.29 |
1 |
4 |
0 |
49 |
299.399 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
65 |
0.48 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
65 |
0.48 |
Binding ≤ 10μM
|
|
CP1A2-1-E |
Cytochrome P450 1A2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2000 |
0.38 |
ADME/T ≤ 10μM
|
|
CP3A4-2-E |
Cytochrome P450 3A4 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
9000 |
0.34 |
ADME/T ≤ 10μM
|
|
Z50590-1-O |
Sus Scrofa (cluster #1 Of 1), Other |
Other |
13 |
0.53 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.33 |
9.9 |
-17.91 |
0 |
4 |
0 |
50 |
277.327 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
3.33 |
10.35 |
-35.25 |
1 |
4 |
1 |
52 |
278.335 |
4 |
↓
|
|
|
|
Analogs
-
3830986
-
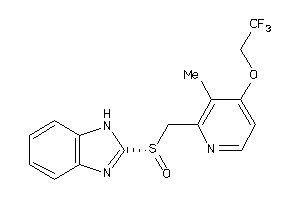
Draw
Identity
99%
90%
80%
70%
Vendors
And 38 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
398 |
0.36 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
398 |
0.36 |
Binding ≤ 10μM
|
|
Z50425-3-O |
Plasmodium Falciparum (cluster #3 Of 22), Other |
Other |
10000 |
0.28 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.88 |
5.41 |
-43.96 |
2 |
5 |
1 |
69 |
370.376 |
6 |
↓
|
|
|
|
Analogs
-
599734
-
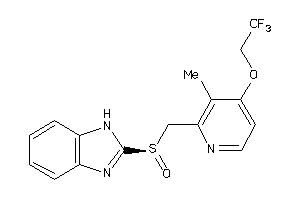
Draw
Identity
99%
90%
80%
70%
Vendors
And 35 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ATP4A-1-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
398 |
0.36 |
Binding ≤ 10μM
|
|
ATP4B-1-E |
Potassium-transporting ATPase Beta Chain (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
398 |
0.36 |
Binding ≤ 10μM
|
|
Z50425-3-O |
Plasmodium Falciparum (cluster #3 Of 22), Other |
Other |
10000 |
0.28 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.88 |
5.61 |
-62.66 |
2 |
5 |
1 |
69 |
370.376 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 28 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
ACES-1-E |
Acetylcholinesterase (cluster #1 Of 12), Eukaryotic |
Eukaryotes |
650 |
0.41 |
Binding ≤ 10μM
|
|
HRH2-1-E |
Histamine H2 Receptor (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
501 |
0.42 |
Binding ≤ 10μM
|
|
ATP4A-2-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
40 |
0.49 |
Functional ≤ 10μM
|
|
ATP4B-2-E |
Potassium-transporting ATPase Beta Chain (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
40 |
0.49 |
Functional ≤ 10μM
|
|
HRH2-2-E |
Histamine H2 Receptor (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
3400 |
0.36 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 27 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
HRH2-2-E |
Histamine H2 Receptor (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
501 |
0.52 |
Binding ≤ 10μM
|
|
ATP4A-2-E |
Potassium-transporting ATPase Alpha Chain 1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
820 |
0.50 |
Functional ≤ 10μM
|
|
ATP4B-2-E |
Potassium-transporting ATPase Beta Chain (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
820 |
0.50 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.14 |
8.62 |
-36.92 |
4 |
6 |
1 |
90 |
253.355 |
7 |
↓
|
|
Ref
Reference (pH 7)
|
0.14 |
7.63 |
-35.87 |
4 |
6 |
1 |
90 |
253.355 |
7 |
↓
|
|
Ref
Reference (pH 7)
|
0.14 |
7.9 |
-14.24 |
3 |
6 |
0 |
89 |
252.347 |
7 |
↓
|
|