|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 46 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
300 |
1.14 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.01 |
-1.64 |
-65.42 |
4 |
4 |
0 |
88 |
119.12 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.01 |
-2.01 |
-46.57 |
3 |
4 |
-1 |
86 |
118.112 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 42 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRG1-6-E |
GABA Receptor Gamma-1 Subunit (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
400 |
1.12 |
Binding ≤ 10μM
|
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
900 |
1.06 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.01 |
-1.68 |
-65.44 |
4 |
4 |
0 |
88 |
119.12 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.01 |
-2.13 |
-46.47 |
3 |
4 |
-1 |
86 |
118.112 |
3 |
↓
|
|
|
|
Analogs
-
388118
-
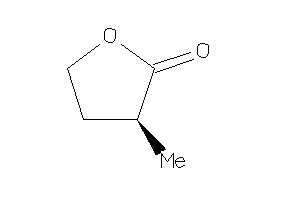
Draw
Identity
99%
90%
80%
70%
Vendors
And 8 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
9200 |
0.88 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.32 |
3.93 |
-7.93 |
0 |
2 |
0 |
26 |
114.144 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 11 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
490 |
0.98 |
Binding ≤ 10μM |
|
Z104301-2-O |
GABA-A Receptor; Anion Channel (cluster #2 Of 2), Other |
Other |
3100 |
0.86 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.42 |
2.86 |
-67.39 |
2 |
3 |
0 |
57 |
127.143 |
1 |
↓
|
|
|
|
Analogs
-
85733
-
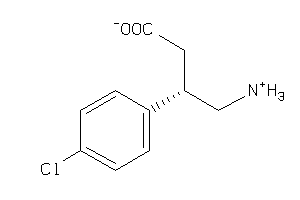
Draw
Identity
99%
90%
80%
70%
Vendors
And 64 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
AA3R-3-E |
Adenosine Receptor A3 (cluster #3 Of 6), Eukaryotic |
Eukaryotes |
50 |
0.73 |
Binding ≤ 10μM
|
|
GABR1-1-E |
GABA-B Receptor 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
32 |
0.75 |
Binding ≤ 10μM
|
|
GABR1-1-E |
GABA-B Receptor 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
60 |
0.72 |
Binding ≤ 10μM
|
|
GABR2-1-E |
GABA-B Receptor 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
32 |
0.75 |
Binding ≤ 10μM
|
|
GABR2-1-E |
GABA-B Receptor 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
60 |
0.72 |
Binding ≤ 10μM
|
|
GBRB1-3-E |
GABA Receptor Beta-1 Subunit (cluster #3 Of 6), Eukaryotic |
Eukaryotes |
330 |
0.65 |
Binding ≤ 10μM
|
|
GBRB2-4-E |
GABA Receptor Beta-2 Subunit (cluster #4 Of 7), Eukaryotic |
Eukaryotes |
330 |
0.65 |
Binding ≤ 10μM
|
|
GBRB3-4-E |
GABA Receptor Beta-3 Subunit (cluster #4 Of 6), Eukaryotic |
Eukaryotes |
330 |
0.65 |
Binding ≤ 10μM
|
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
30 |
0.75 |
Binding ≤ 10μM
|
|
Z50512-6-O |
Cavia Porcellus (cluster #6 Of 7), Other |
Other |
6400 |
0.52 |
Functional ≤ 10μM |
|
Z50597-12-O |
Rattus Norvegicus (cluster #12 Of 12), Other |
Other |
330 |
0.65 |
Functional ≤ 10μM
|
|
Z50597-12-O |
Rattus Norvegicus (cluster #12 Of 12), Other |
Other |
1800 |
0.57 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.42 |
4.81 |
-48.33 |
3 |
3 |
0 |
68 |
213.664 |
4 |
↓
|
|
|
|
Analogs
-
61
-
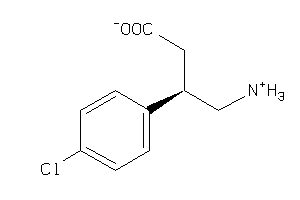
Draw
Identity
99%
90%
80%
70%
Vendors
And 53 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.42 |
4.85 |
-63 |
3 |
3 |
0 |
68 |
213.664 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
500 |
0.88 |
Binding ≤ 10μM |
|
Z104301-2-O |
GABA-A Receptor; Anion Channel (cluster #2 Of 2), Other |
Other |
3400 |
0.77 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.48 |
2.17 |
-35.89 |
3 |
4 |
0 |
80 |
160.198 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
500 |
0.88 |
Binding ≤ 10μM |
|
Z104301-2-O |
GABA-A Receptor; Anion Channel (cluster #2 Of 2), Other |
Other |
3400 |
0.77 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.48 |
2.17 |
-35.89 |
3 |
4 |
0 |
80 |
160.198 |
2 |
↓
|
|
|
|
Analogs
-
13734075
-
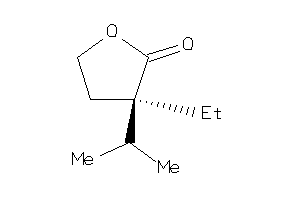
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
35 |
0.95 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.23 |
5.34 |
-7.77 |
0 |
2 |
0 |
26 |
156.225 |
2 |
↓
|
|
|
|
Analogs
-
13734071
-
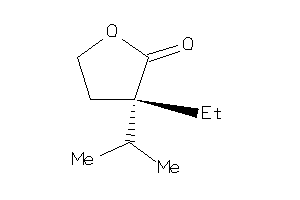
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z104301-5-O |
GABA-A Receptor; Anion Channel (cluster #5 Of 8), Other |
Other |
35 |
0.95 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
1.23 |
5.35 |
-8.33 |
0 |
2 |
0 |
26 |
156.225 |
2 |
↓
|
|