|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CNR1-3-E |
Cannabinoid CB1 Receptor (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
4 |
0.47 |
Binding ≤ 10μM
|
|
CNR2-2-E |
Cannabinoid CB2 Receptor (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
195 |
0.38 |
Binding ≤ 10μM
|
|
FAAH1-6-E |
Anandamide Amidohydrolase (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
900 |
0.34 |
Binding ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
420 |
0.36 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
6.88 |
13.99 |
-9.23 |
1 |
2 |
0 |
29 |
365.989 |
16 |
↓
|
|
|
|
Analogs
-
4583060
-
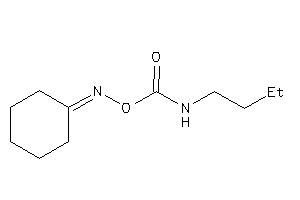
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
DGLA-1-E |
Sn1-specific Diacylglycerol Lipase Alpha (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
60 |
0.36 |
Binding ≤ 10μM
|
|
DGLB-1-E |
Sn1-specific Diacylglycerol Lipase Beta (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
60 |
0.36 |
Binding ≤ 10μM
|
|
NCEH1-1-E |
Neutral Cholesterol Ester Hydrolase 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
10000 |
0.25 |
Binding ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
1000 |
0.30 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.79 |
8.43 |
-20.19 |
2 |
8 |
0 |
101 |
394.516 |
11 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
5000 |
0.82 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.90 |
-1.85 |
-50.71 |
3 |
4 |
1 |
57 |
129.139 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
-0.90 |
-3.11 |
-12.29 |
2 |
4 |
0 |
53 |
128.131 |
0 |
↓
|
|
Mid
Mid (pH 6-8)
|
-0.90 |
-0.82 |
-69.59 |
2 |
4 |
0 |
60 |
128.131 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 25 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
80 |
0.66 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
5500 |
0.49 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
7600 |
0.48 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
320 |
0.61 |
Binding ≤ 10μM
|
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
500 |
0.59 |
Binding ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3000 |
0.52 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.63 |
1.9 |
-54.25 |
6 |
9 |
0 |
164 |
219.201 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.57 |
-53.16 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.63 |
1.57 |
-48.12 |
5 |
9 |
-1 |
162 |
218.193 |
7 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
4000 |
0.16 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.85 |
-4.62 |
-92.12 |
16 |
17 |
2 |
300 |
658.846 |
23 |
↓
|
|
Mid
Mid (pH 6-8)
|
-1.85 |
-4.98 |
-53.19 |
15 |
17 |
1 |
298 |
657.838 |
23 |
↓
|
|
Mid
Mid (pH 6-8)
|
-1.98 |
-4.63 |
-53.02 |
15 |
17 |
1 |
301 |
657.838 |
22 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 8 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
4300 |
0.58 |
Binding ≤ 10μM
|
|
NOS2-7-E |
Nitric Oxide Synthase, Inducible (cluster #7 Of 9), Eukaryotic |
Eukaryotes |
860 |
0.65 |
Binding ≤ 10μM |
|
NOS3-1-E |
Nitric Oxide Synthase, Endothelial (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
4200 |
0.58 |
Binding ≤ 10μM
|
|
NOS1-2-E |
Nitric-oxide Synthase, Brain (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
100 |
0.75 |
Functional ≤ 10μM
|
|
NOS2-1-E |
Nitric Oxide Synthase, Inducible (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Functional ≤ 10μM
|
|
NOS3-1-E |
Nitric-oxide Synthase, Endothelial (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.70 |
Functional ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3000 |
0.59 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.26 |
1.39 |
-72.47 |
7 |
6 |
1 |
117 |
189.239 |
7 |
↓
|
|
Hi
High (pH 8-9.5)
|
8.62 |
17.49 |
-47.16 |
0 |
0 |
-1 |
0 |
421.61 |
5 |
↓
|
|
|
|
Analogs
-
5855933
-
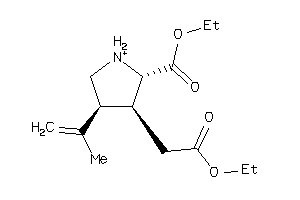
-
34599078
-
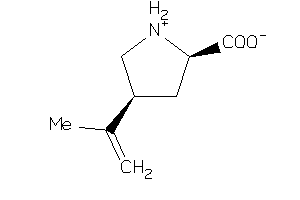
-
1481864
-
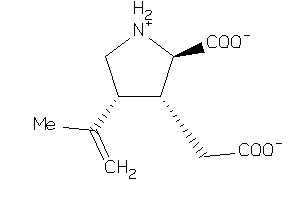
-
2008786
-
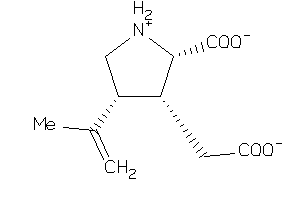
-
3872987
-
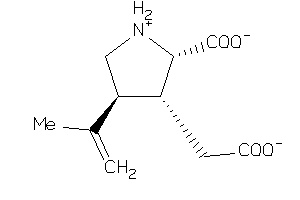
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
AA3R-6-E |
Adenosine Receptor A3 (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
77 |
0.66 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
7500 |
0.48 |
Binding ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.49 |
Binding ≤ 10μM
|
|
GRIA3-3-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.49 |
Binding ≤ 10μM
|
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
1714 |
0.54 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
177 |
0.63 |
Binding ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
53 |
0.68 |
Binding ≤ 10μM
|
|
GRIK3-2-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
10 |
0.75 |
Binding ≤ 10μM
|
|
GRIK4-2-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
7 |
0.76 |
Binding ≤ 10μM
|
|
GRIK5-2-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
7 |
0.76 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
7510 |
0.48 |
Functional ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
700 |
0.57 |
Functional ≤ 10μM
|
|
Z50597-4-O |
Rattus Norvegicus (cluster #4 Of 5), Other |
Other |
18 |
0.72 |
Binding ≤ 10μM
|
|
Z50466-5-O |
Trypanosoma Cruzi (cluster #5 Of 8), Other |
Other |
3000 |
0.52 |
Functional ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3700 |
0.51 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.81 |
5.21 |
-88.22 |
2 |
5 |
-1 |
97 |
212.225 |
4 |
↓
|
|
|
|
Analogs
-
27865868
-
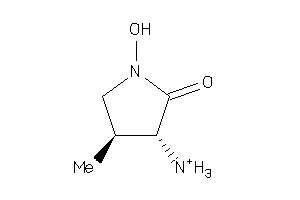
-
34369915
-
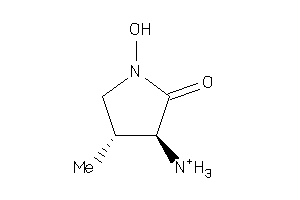
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NMDE3-3-E |
Glutamate [NMDA] Receptor Subunit Epsilon 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
650 |
0.96 |
Binding ≤ 10μM
|
|
NMDZ1-4-E |
Glutamate (NMDA) Receptor Subunit Zeta 1 (cluster #4 Of 6), Eukaryotic |
Eukaryotes |
2900 |
0.86 |
Binding ≤ 10μM |
|
Z104302-3-O |
Glutamate NMDA Receptor (cluster #3 Of 7), Other |
Other |
2700 |
0.87 |
Binding ≤ 10μM |
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
650 |
0.96 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.18 |
-3.93 |
-48.29 |
4 |
4 |
1 |
68 |
131.155 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.18 |
-3.45 |
-68.24 |
3 |
4 |
0 |
71 |
130.147 |
0 |
↓
|
|
|
|
Analogs
-
4468952
-
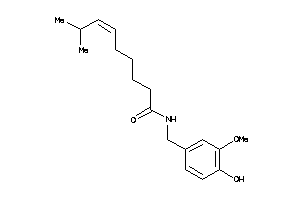
-
33845128
-
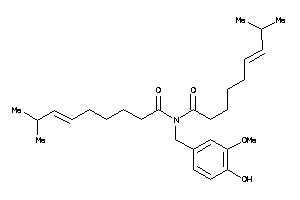
Draw
Identity
99%
90%
80%
70%
Vendors
And 42 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.10 |
6.6 |
-9.89 |
2 |
4 |
0 |
59 |
305.418 |
9 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.69 |
1.88 |
-12.03 |
1 |
3 |
0 |
54 |
326.436 |
6 |
↓
|
|
|
|
Analogs
-
6158191
-

Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIK3-2-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
700 |
0.78 |
Binding ≤ 10μM |
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
5000 |
0.67 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.38 |
-2.22 |
-80.43 |
2 |
5 |
-1 |
97 |
158.133 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 14 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
NMDE3-3-E |
Glutamate [NMDA] Receptor Subunit Epsilon 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
2500 |
0.98 |
Binding ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
6300 |
0.91 |
Functional ≤ 10μM |
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.66 |
-2.73 |
-51.23 |
4 |
4 |
1 |
68 |
117.128 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.66 |
-3.03 |
-8.73 |
3 |
4 |
0 |
67 |
116.12 |
0 |
↓
|
|
Hi
High (pH 8-9.5)
|
-1.66 |
-1.72 |
-69.53 |
3 |
4 |
0 |
71 |
116.12 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
476 |
0.63 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.63 |
2.46 |
-50.7 |
1 |
6 |
-1 |
99 |
219.242 |
5 |
↓
|
|
|
|
|
|
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
492 |
0.63 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-1.63 |
2.51 |
-49.85 |
1 |
6 |
-1 |
99 |
219.242 |
5 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 14 More
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.42 |
2.83 |
-58.8 |
2 |
3 |
0 |
57 |
127.143 |
1 |
↓
|
|
|
|
Analogs
-
3873971
-
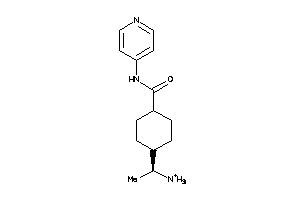
Draw
Identity
99%
90%
80%
70%
Vendors
And 20 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
CCL2-1-E |
C-C Motif Chemokine 2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
2630 |
0.43 |
Binding ≤ 10μM
|
|
KPCE-1-E |
Protein Kinase C Epsilon (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
335 |
0.50 |
Binding ≤ 10μM
|
|
KS6A5-2-E |
Ribosomal Protein S6 Kinase Alpha 5 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
8300 |
0.40 |
Binding ≤ 10μM
|
|
PKN2-1-E |
Protein Kinase N2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
70 |
0.56 |
Binding ≤ 10μM
|
|
PKN2-1-E |
Protein Kinase N2 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
600 |
0.48 |
Binding ≤ 10μM
|
|
ROCK1-2-E |
Rho-associated Protein Kinase 1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
140 |
0.53 |
Binding ≤ 10μM
|
|
ROCK1-2-E |
Rho-associated Protein Kinase 1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
871 |
0.47 |
Binding ≤ 10μM
|
|
ROCK2-2-E |
Rho-associated Protein Kinase 2 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
245 |
0.51 |
Binding ≤ 10μM
|
|
ROCK2-2-E |
Rho-associated Protein Kinase 2 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
800 |
0.47 |
Binding ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
4750 |
0.41 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.72 |
3.52 |
-39.49 |
4 |
4 |
1 |
70 |
248.35 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
1.72 |
5.1 |
-91.09 |
7 |
8 |
2 |
122 |
438.507 |
5 |
↓
|
|
|
|
|