Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.67 |
10.61 |
-8.96 |
1 |
5 |
0 |
56 |
414.292 |
2 |
↓
|
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
PDE5A-1-E |
Phosphodiesterase 5A (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
8750 |
0.25 |
Binding ≤ 10μM
|
|
Z81252-3-O |
MDA-MB-231 (Breast Adenocarcinoma Cells) (cluster #3 Of 11), Other |
Other |
4780 |
0.27 |
Functional ≤ 10μM
|
Reactome Annotations from Targets (via Uniprot)
| Description |
Species |
|
cGMP effects |
|
Rings
-
Pyrrole
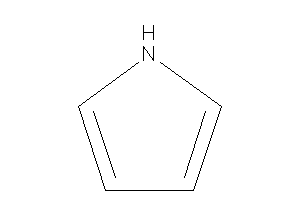
-
1,2,3,6-tetrahydropyridine
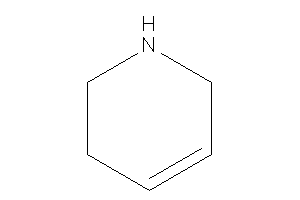
-
Benzene
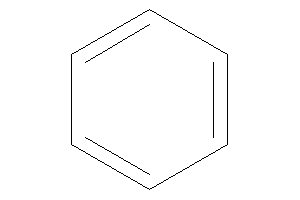
-
Hydantoin
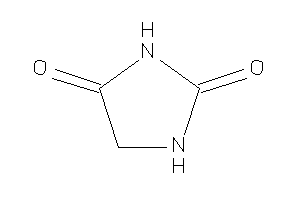
-
3a,4,9,10-tetrahydroimidazo[1,5-…
![3a,4,9,10-tetrahydroimidazo[1,5-b]$b-carboline-1,3-quinone](//zinc12.docking.org/img/rings/231241.gif)
-
10-phenyl-3a,4,9,10-tetrahydroim…
![10-phenyl-3a,4,9,10-tetrahydroimidazo[1,5-b]$b-carboline-1,3-quinone](//zinc12.docking.org/img/rings/231846.gif)
No pre-computed analogs available. Try a structural similarity search.