|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.22 |
7.07 |
-10.5 |
2 |
3 |
0 |
44 |
257.296 |
0 |
↓
|
|
Lo
Low (pH 4.5-6)
|
4.22 |
7.33 |
-31.17 |
3 |
3 |
1 |
46 |
258.304 |
0 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
1 |
0.63 |
Binding ≤ 10μM
|
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
1 |
0.63 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
15 |
0.55 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
15 |
0.55 |
Binding ≤ 10μM
|
|
GBRA3-7-E |
GABA Receptor Alpha-3 Subunit (cluster #7 Of 8), Eukaryotic |
Eukaryotes |
19 |
0.54 |
Binding ≤ 10μM
|
|
GBRA3-7-E |
GABA Receptor Alpha-3 Subunit (cluster #7 Of 8), Eukaryotic |
Eukaryotes |
19 |
0.54 |
Binding ≤ 10μM
|
|
GBRA4-1-E |
GABA Receptor Alpha-4 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
1000 |
0.42 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
111 |
0.49 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
111 |
0.49 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
19 |
0.54 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
19 |
0.54 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
19 |
0.54 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
19 |
0.54 |
Binding ≤ 10μM
|
|
Z104301-1-O |
GABA-A Receptor; Anion Channel (cluster #1 Of 8), Other |
Other |
10 |
0.56 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.85 |
0.6 |
-9.17 |
1 |
4 |
0 |
54 |
268.316 |
3 |
↓
|
|
|
|
Analogs
-
26507557
-
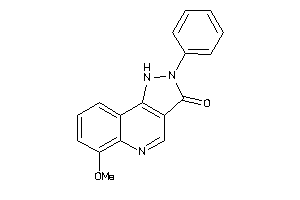
-
13741323
-
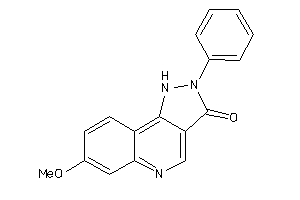
Draw
Identity
99%
90%
80%
70%
Vendors
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.34 |
4.47 |
-12.66 |
1 |
5 |
0 |
60 |
291.31 |
2 |
↓
|
|
Ref
Reference (pH 7)
|
2.88 |
6.29 |
-11.45 |
1 |
5 |
0 |
60 |
291.31 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 7 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
1 |
0.70 |
Binding ≤ 10μM
|
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
5 |
0.65 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
5 |
0.65 |
Binding ≤ 10μM
|
|
GBRA3-7-E |
GABA Receptor Alpha-3 Subunit (cluster #7 Of 8), Eukaryotic |
Eukaryotes |
6 |
0.64 |
Binding ≤ 10μM
|
|
GBRA3-7-E |
GABA Receptor Alpha-3 Subunit (cluster #7 Of 8), Eukaryotic |
Eukaryotes |
6 |
0.64 |
Binding ≤ 10μM
|
|
GBRA4-1-E |
GABA Receptor Alpha-4 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
27 |
0.59 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
27 |
0.59 |
Binding ≤ 10μM
|
|
GBRA6-8-E |
GABA Receptor Alpha-6 Subunit (cluster #8 Of 8), Eukaryotic |
Eukaryotes |
2700 |
0.43 |
Binding ≤ 10μM
|
|
GBRA6-8-E |
GABA Receptor Alpha-6 Subunit (cluster #8 Of 8), Eukaryotic |
Eukaryotes |
2700 |
0.43 |
Binding ≤ 10μM
|
|
GBRB1-1-E |
GABA Receptor Beta-1 Subunit (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRB2-6-E |
GABA Receptor Beta-2 Subunit (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
6 |
0.64 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRD-5-E |
GABA Receptor Delta Subunit (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRE-5-E |
GABA Receptor Epsilon Subunit (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRG1-1-E |
GABA Receptor Gamma-1 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
1 |
0.70 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
6 |
0.64 |
Binding ≤ 10μM
|
|
GBRP-5-E |
GABA Receptor Pi Subunit (cluster #5 Of 6), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRR1-1-E |
GABA Receptor Rho-1 Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
GBRR2-1-E |
GABA Receptor Rho-2 Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
PDE5A-1-E |
Phosphodiesterase 5A (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
800 |
0.47 |
Binding ≤ 10μM
|
|
Q91ZM7-1-E |
GABA Receptor Theta Subunit (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
7 |
0.63 |
Binding ≤ 10μM
|
|
Z104301-1-O |
GABA-A Receptor; Anion Channel (cluster #1 Of 8), Other |
Other |
5 |
0.65 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.04 |
6.49 |
-9.28 |
1 |
4 |
0 |
55 |
240.262 |
3 |
↓
|
|
Mid
Mid (pH 6-8)
|
3.04 |
6.94 |
-33.36 |
2 |
4 |
1 |
56 |
241.27 |
3 |
↓
|
|
|
|
|
|
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
1 |
0.74 |
Binding ≤ 10μM
|
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
2 |
0.72 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRA3-7-E |
GABA Receptor Alpha-3 Subunit (cluster #7 Of 8), Eukaryotic |
Eukaryotes |
5 |
0.68 |
Binding ≤ 10μM
|
|
GBRA4-1-E |
GABA Receptor Alpha-4 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
7 |
0.67 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
5 |
0.68 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
58 |
0.60 |
Binding ≤ 10μM
|
|
GBRA6-8-E |
GABA Receptor Alpha-6 Subunit (cluster #8 Of 8), Eukaryotic |
Eukaryotes |
8 |
0.67 |
Binding ≤ 10μM
|
|
GBRB1-1-E |
GABA Receptor Beta-1 Subunit (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRB2-6-E |
GABA Receptor Beta-2 Subunit (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
8 |
0.67 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRD-5-E |
GABA Receptor Delta Subunit (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRE-5-E |
GABA Receptor Epsilon Subunit (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRG1-1-E |
GABA Receptor Gamma-1 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
3 |
0.70 |
Binding ≤ 10μM
|
|
GBRP-5-E |
GABA Receptor Pi Subunit (cluster #5 Of 6), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRR1-1-E |
GABA Receptor Rho-1 Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
GBRR2-1-E |
GABA Receptor Rho-2 Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
Q91ZM7-1-E |
GABA Receptor Theta Subunit (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4 |
0.69 |
Binding ≤ 10μM
|
|
Z104301-1-O |
GABA-A Receptor; Anion Channel (cluster #1 Of 8), Other |
Other |
5 |
0.68 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.66 |
5.56 |
-9.44 |
1 |
4 |
0 |
55 |
226.235 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.66 |
6.01 |
-33.94 |
2 |
4 |
1 |
56 |
227.243 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 4 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
17 |
0.49 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
81 |
0.45 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
1445 |
0.37 |
Binding ≤ 10μM
|
|
GBRB2-6-E |
GABA Receptor Beta-2 Subunit (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
81 |
0.45 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
1445 |
0.37 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
81 |
0.45 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.92 |
-2.17 |
-8.96 |
2 |
4 |
0 |
61 |
357.207 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRA3-7-E |
GABA Receptor Alpha-3 Subunit (cluster #7 Of 8), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRA4-1-E |
GABA Receptor Alpha-4 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRA6-8-E |
GABA Receptor Alpha-6 Subunit (cluster #8 Of 8), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRB1-1-E |
GABA Receptor Beta-1 Subunit (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRB2-6-E |
GABA Receptor Beta-2 Subunit (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRD-5-E |
GABA Receptor Delta Subunit (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRE-5-E |
GABA Receptor Epsilon Subunit (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRG1-1-E |
GABA Receptor Gamma-1 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRP-5-E |
GABA Receptor Pi Subunit (cluster #5 Of 6), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRR1-1-E |
GABA Receptor Rho-1 Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
GBRR2-1-E |
GABA Receptor Rho-2 Subunit (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
|
Q91ZM7-1-E |
GABA Receptor Theta Subunit (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
197 |
0.55 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.02 |
3.27 |
-10.64 |
2 |
4 |
0 |
58 |
225.251 |
1 |
↓
|
|
Mid
Mid (pH 6-8)
|
2.02 |
3.76 |
-38.44 |
3 |
4 |
1 |
59 |
226.259 |
1 |
↓
|
|
|
|
Analogs
-
4163159
-
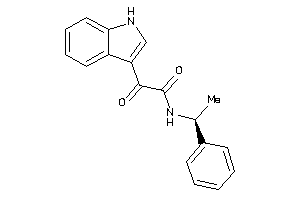
Draw
Identity
99%
90%
80%
70%
Vendors
And 2 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRA1-6-E |
GABA Receptor Alpha-1 Subunit (cluster #6 Of 8), Eukaryotic |
Eukaryotes |
1150 |
0.38 |
Binding ≤ 10μM
|
|
GBRA2-5-E |
GABA Receptor Alpha-2 Subunit (cluster #5 Of 8), Eukaryotic |
Eukaryotes |
1550 |
0.37 |
Binding ≤ 10μM
|
|
GBRA3-7-E |
GABA Receptor Alpha-3 Subunit (cluster #7 Of 8), Eukaryotic |
Eukaryotes |
1307 |
0.37 |
Binding ≤ 10μM
|
|
GBRA4-1-E |
GABA Receptor Alpha-4 Subunit (cluster #1 Of 7), Eukaryotic |
Eukaryotes |
1307 |
0.37 |
Binding ≤ 10μM
|
|
GBRA5-3-E |
GABA Receptor Alpha-5 Subunit (cluster #3 Of 8), Eukaryotic |
Eukaryotes |
5500 |
0.33 |
Binding ≤ 10μM
|
|
GBRB1-1-E |
GABA Receptor Beta-1 Subunit (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
1307 |
0.37 |
Binding ≤ 10μM
|
|
GBRB2-6-E |
GABA Receptor Beta-2 Subunit (cluster #6 Of 7), Eukaryotic |
Eukaryotes |
1150 |
0.38 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
5500 |
0.33 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
1307 |
0.37 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
2.67 |
-0.65 |
-8.81 |
2 |
4 |
0 |
61 |
292.338 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GBRA1-1-E |
GABA Receptor Alpha-1 Subunit (cluster #1 Of 8), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
GBRA2-1-E |
GABA Receptor Alpha-2 Subunit (cluster #1 Of 8), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
GBRA3-1-E |
GABA Receptor Alpha-3 Subunit (cluster #1 Of 8), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
GBRA5-1-E |
GABA Receptor Alpha-5 Subunit (cluster #1 Of 8), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
GBRA6-8-E |
GABA Receptor Alpha-6 Subunit (cluster #8 Of 8), Eukaryotic |
Eukaryotes |
4 |
0.56 |
Binding ≤ 10μM
|
|
GBRB3-2-E |
GABA Receptor Beta-3 Subunit (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
4 |
0.56 |
Binding ≤ 10μM
|
|
GBRG2-2-E |
GABA Receptor Gamma-2 Subunit (cluster #2 Of 7), Eukaryotic |
Eukaryotes |
4 |
0.56 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.09 |
5.83 |
-11.18 |
1 |
4 |
0 |
51 |
340.18 |
1 |
↓
|
|
Ref
Reference (pH 7)
|
3.63 |
7.59 |
-9.82 |
1 |
4 |
0 |
51 |
340.18 |
1 |
↓
|
|