|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
120 |
0.57 |
Binding ≤ 10μM
|
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
279 |
0.54 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
115 |
0.57 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
120 |
0.57 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
120 |
0.57 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
120 |
0.57 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
207 |
0.55 |
Binding ≤ 10μM
|
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
3700 |
0.45 |
Binding ≤ 10μM
|
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3700 |
0.45 |
Binding ≤ 10μM
|
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3700 |
0.45 |
Binding ≤ 10μM
|
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3700 |
0.45 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
393 |
0.53 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3700 |
0.45 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.61 |
-1.5 |
-58.63 |
3 |
10 |
-1 |
172 |
243.155 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.61 |
-1.72 |
-91.85 |
2 |
10 |
-2 |
170 |
242.147 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
65 |
0.67 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
53 |
0.68 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
451 |
0.59 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
451 |
0.59 |
Binding ≤ 10μM
|
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
47 |
0.68 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
57 |
0.68 |
Binding ≤ 10μM
|
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
2110 |
0.53 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.35 |
-0.77 |
-39.5 |
4 |
7 |
0 |
123 |
233.611 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
163 |
0.63 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
176 |
0.63 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
976 |
0.56 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
972 |
0.56 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
2 |
0.81 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
0 |
0.00 |
Binding ≤ 10μM
|
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
140 |
0.64 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.95 |
0.82 |
-42.22 |
4 |
7 |
0 |
123 |
325.062 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
-2.49 |
-1.87 |
-67.59 |
3 |
7 |
-1 |
126 |
324.054 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 5 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
386 |
0.64 |
Binding ≤ 10μM
|
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
1800 |
0.57 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
898 |
0.60 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
1800 |
0.57 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
1800 |
0.57 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
1800 |
0.57 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
8850 |
0.51 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-4.25 |
-0.27 |
-47.38 |
4 |
7 |
0 |
123 |
199.166 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.82 |
-3.19 |
-102.6 |
2 |
7 |
-2 |
124 |
197.15 |
3 |
↓
|
|
Hi
High (pH 8-9.5)
|
-3.82 |
-2.83 |
-75.1 |
3 |
7 |
-1 |
126 |
198.158 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 2 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
15 |
0.73 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
25 |
0.71 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
70 |
0.67 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
305 |
0.61 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
1380 |
0.55 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1820 |
0.54 |
Binding ≤ 10μM
|
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
382 |
0.60 |
Functional ≤ 10μM
|
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
463 |
0.59 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.87 |
-0.19 |
-45.44 |
4 |
7 |
0 |
123 |
217.156 |
3 |
↓
|
|
Mid
Mid (pH 6-8)
|
-3.41 |
-2.93 |
-72.07 |
3 |
7 |
-1 |
126 |
216.148 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 7 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
360 |
0.50 |
Binding ≤ 10μM
|
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
360 |
0.50 |
Binding ≤ 10μM
|
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
360 |
0.50 |
Binding ≤ 10μM
|
|
GRIA4-2-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
360 |
0.50 |
Binding ≤ 10μM
|
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
2900 |
0.43 |
Binding ≤ 10μM |
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2900 |
0.43 |
Binding ≤ 10μM |
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2900 |
0.43 |
Binding ≤ 10μM |
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2900 |
0.43 |
Binding ≤ 10μM |
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
740 |
0.48 |
Binding ≤ 10μM
|
|
NMD3A-4-E |
Glutamate [NMDA] Receptor Subunit 3A (cluster #4 Of 6), Eukaryotic |
Eukaryotes |
390 |
0.50 |
Binding ≤ 10μM
|
|
NMD3B-4-E |
Glutamate [NMDA] Receptor Subunit 3B (cluster #4 Of 6), Eukaryotic |
Eukaryotes |
390 |
0.50 |
Binding ≤ 10μM
|
|
NMDE1-1-E |
Glutamate [NMDA] Receptor Subunit Epsilon 1 (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
840 |
0.47 |
Binding ≤ 10μM |
|
NMDE2-1-E |
Glutamate [NMDA] Receptor Subunit Epsilon 2 (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
840 |
0.47 |
Binding ≤ 10μM |
|
NMDE3-2-E |
Glutamate [NMDA] Receptor Subunit Epsilon 3 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
840 |
0.47 |
Binding ≤ 10μM |
|
NMDE4-4-E |
Glutamate [NMDA] Receptor Subunit Epsilon 4 (cluster #4 Of 6), Eukaryotic |
Eukaryotes |
390 |
0.50 |
Binding ≤ 10μM
|
|
NMDZ1-1-E |
Glutamate (NMDA) Receptor Subunit Zeta 1 (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
840 |
0.47 |
Binding ≤ 10μM |
|
Z104302-5-O |
Glutamate NMDA Receptor (cluster #5 Of 7), Other |
Other |
4500 |
0.42 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.46 |
1.23 |
-35.53 |
1 |
10 |
-1 |
160 |
251.134 |
2 |
↓
|
|
Ref
Reference (pH 7)
|
0.42 |
3.67 |
-137.39 |
2 |
10 |
2 |
154 |
252.142 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.42 |
2.62 |
-51.15 |
1 |
10 |
1 |
153 |
251.134 |
2 |
↓
|
|
|
|
Analogs
-
2311806
-
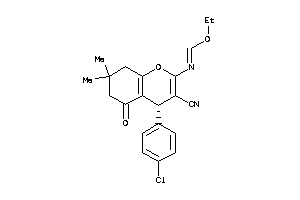
-
2466126
-
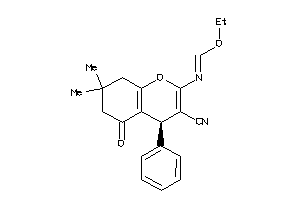
-
2466129
-
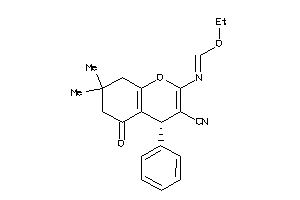
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1600 |
0.31 |
Binding ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
1600 |
0.31 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.26 |
1.39 |
-15.86 |
2 |
4 |
0 |
76 |
344.414 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1600 |
0.31 |
Binding ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
1600 |
0.31 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.26 |
1.39 |
-15.8 |
2 |
4 |
0 |
76 |
344.414 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIA4-2-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
135 |
0.48 |
Binding ≤ 10μM |
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM |
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2200 |
0.40 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
260 |
0.46 |
Functional ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
300 |
0.46 |
Functional ≤ 10μM
|
|
Z104302-4-O |
Glutamate NMDA Receptor (cluster #4 Of 7), Other |
Other |
7400 |
0.36 |
Binding ≤ 10μM |
|
Z50574-1-O |
Xenopus Sp. (cluster #1 Of 2), Other |
Other |
260 |
0.46 |
Functional ≤ 10μM
|
|
Z50597-8-O |
Rattus Norvegicus (cluster #8 Of 12), Other |
Other |
810 |
0.43 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.26 |
2.06 |
-40.89 |
1 |
9 |
-1 |
132 |
272.2 |
2 |
↓
|
|
Ref
Reference (pH 7)
|
0.22 |
5.15 |
-137.07 |
2 |
9 |
2 |
126 |
273.208 |
2 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.22 |
4.1 |
-49.04 |
1 |
9 |
1 |
125 |
272.2 |
2 |
↓
|
|
|
|
Analogs
-
19850502
-
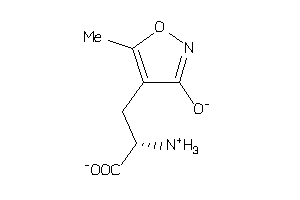
Draw
Identity
99%
90%
80%
70%
Vendors
And 20 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
364 |
0.69 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.57 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
364 |
0.69 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.57 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
40 |
0.80 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.57 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
40 |
0.80 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
4800 |
0.57 |
Binding ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
740 |
0.66 |
Binding ≤ 10μM
|
|
GRIK2-3-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIK3-3-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3310 |
0.59 |
Functional ≤ 10μM
|
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
3310 |
0.59 |
Functional ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
800 |
0.66 |
Functional ≤ 10μM
|
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
4800 |
0.57 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.69 |
-1.91 |
-52.1 |
3 |
6 |
-1 |
117 |
185.159 |
3 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-2.69 |
-2.52 |
-37.61 |
4 |
6 |
0 |
114 |
186.167 |
3 |
↓
|
|
|
|
Analogs
-
19850499
-
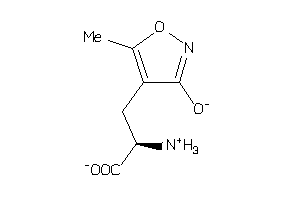
Draw
Identity
99%
90%
80%
70%
Vendors
And 23 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
22 |
0.82 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
364 |
0.69 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
17 |
0.84 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
364 |
0.69 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
21 |
0.83 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
40 |
0.80 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
40 |
0.80 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
40 |
0.80 |
Binding ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
740 |
0.66 |
Binding ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
1150 |
0.64 |
Binding ≤ 10μM
|
|
GRIK2-3-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIK3-3-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
12 |
0.85 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2144 |
0.61 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3310 |
0.59 |
Functional ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
3310 |
0.59 |
Functional ≤ 10μM
|
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
3500 |
0.59 |
Functional ≤ 10μM
|
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
3800 |
0.58 |
Functional ≤ 10μM
|
|
Z104302-1-O |
Glutamate NMDA Receptor (cluster #1 Of 7), Other |
Other |
3500 |
0.59 |
Binding ≤ 10μM
|
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
800 |
0.66 |
Functional ≤ 10μM
|
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
6700 |
0.56 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-2.69 |
-1.72 |
-76.52 |
3 |
6 |
-1 |
117 |
185.159 |
3 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
9200 |
0.35 |
Binding ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM
|
|
GRIA3-3-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM
|
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
4800 |
0.37 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
4200 |
0.38 |
Binding ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
2100 |
0.40 |
Binding ≤ 10μM
|
|
GRIK4-2-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
4200 |
0.38 |
Binding ≤ 10μM
|
|
GRIK5-2-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
4200 |
0.38 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Functional ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Functional ≤ 10μM
|
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Functional ≤ 10μM
|
|
GRIA4-2-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.37 |
Functional ≤ 10μM
|
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
230 |
0.46 |
Functional ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
230 |
0.46 |
Functional ≤ 10μM
|
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
230 |
0.46 |
Functional ≤ 10μM
|
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
230 |
0.46 |
Functional ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
230 |
0.46 |
Functional ≤ 10μM
|
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
6000 |
0.37 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.53 |
4.85 |
-68.53 |
2 |
7 |
-1 |
110 |
278.336 |
4 |
↓
|
|
Mid
Mid (pH 6-8)
|
-0.53 |
3.91 |
-46.21 |
3 |
7 |
0 |
111 |
279.344 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 8 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
70 |
0.44 |
Binding ≤ 10μM |
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
70 |
0.44 |
Binding ≤ 10μM |
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
70 |
0.44 |
Binding ≤ 10μM |
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 5), Eukaryotic |
Eukaryotes |
70 |
0.44 |
Binding ≤ 10μM |
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
8000 |
0.31 |
Binding ≤ 10μM |
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
8000 |
0.31 |
Binding ≤ 10μM |
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
8000 |
0.31 |
Binding ≤ 10μM |
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
8000 |
0.31 |
Binding ≤ 10μM |
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
79 |
0.43 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
6000 |
0.32 |
Functional ≤ 10μM
|
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
2500 |
0.34 |
Functional ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
200 |
0.41 |
Functional ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
200 |
0.41 |
Functional ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
90 |
0.43 |
Functional ≤ 10μM
|
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
90 |
0.43 |
Functional ≤ 10μM
|
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
90 |
0.43 |
Functional ≤ 10μM
|
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
90 |
0.43 |
Functional ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
90 |
0.43 |
Functional ≤ 10μM
|
|
Z104302-4-O |
Glutamate NMDA Receptor (cluster #4 Of 7), Other |
Other |
200 |
0.41 |
Binding ≤ 10μM
|
|
Z50594-1-O |
Mus Musculus (cluster #1 Of 9), Other |
Other |
300 |
0.40 |
Functional ≤ 10μM |
|
Z50597-1-O |
Rattus Norvegicus (cluster #1 Of 12), Other |
Other |
4000 |
0.33 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.57 |
1.41 |
-25.8 |
4 |
10 |
0 |
172 |
336.285 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
0.33 |
-0.74 |
-49.41 |
3 |
10 |
-1 |
175 |
335.277 |
2 |
↓
|
|
Mid
Mid (pH 6-8)
|
0.33 |
-0.78 |
-57.47 |
3 |
10 |
-1 |
175 |
335.277 |
2 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 36 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
2000 |
0.53 |
Binding ≤ 10μM |
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
2000 |
0.53 |
Binding ≤ 10μM |
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
2000 |
0.53 |
Binding ≤ 10μM |
|
GRIA4-2-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
2000 |
0.53 |
Binding ≤ 10μM |
|
Z104302-4-O |
Glutamate NMDA Receptor (cluster #4 Of 7), Other |
Other |
8300 |
0.47 |
Binding ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.33 |
2.11 |
-16.25 |
2 |
7 |
0 |
112 |
207.145 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.57 |
1.05 |
-41.16 |
1 |
7 |
-1 |
115 |
206.137 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 3 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
2100 |
0.50 |
Binding ≤ 10μM
|
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
2780 |
0.49 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
1800 |
0.50 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
2100 |
0.50 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
2100 |
0.50 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
7900 |
0.45 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
2100 |
0.50 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
7600 |
0.45 |
Binding ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2 |
0.76 |
Binding ≤ 10μM
|
|
GRIK3-3-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
2530 |
0.49 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
480 |
0.55 |
Binding ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
660 |
0.54 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.97 |
-0.38 |
-49.09 |
3 |
6 |
-1 |
117 |
227.24 |
4 |
↓
|
|
Hi
High (pH 8-9.5)
|
-0.97 |
-0.69 |
-105.71 |
2 |
6 |
-2 |
115 |
226.232 |
4 |
↓
|
|
Lo
Low (pH 4.5-6)
|
-1.42 |
1.24 |
-28.43 |
4 |
6 |
0 |
114 |
228.248 |
4 |
↓
|
|
|
|
Analogs
-
5855933
-
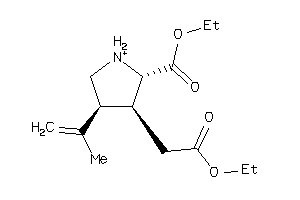
-
34599078
-
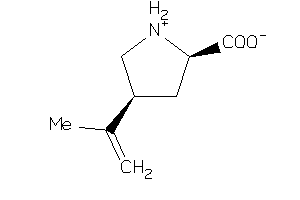
-
1481864
-
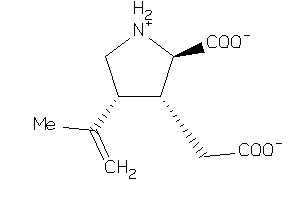
-
2008786
-
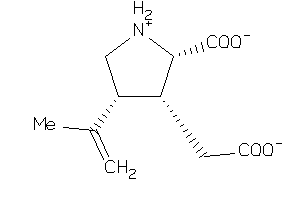
-
3872987
-
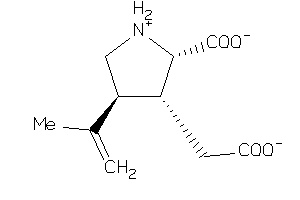
Draw
Identity
99%
90%
80%
70%
Vendors
And 9 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
AA3R-6-E |
Adenosine Receptor A3 (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
77 |
0.66 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
7500 |
0.48 |
Binding ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.49 |
Binding ≤ 10μM
|
|
GRIA3-3-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6000 |
0.49 |
Binding ≤ 10μM
|
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
1714 |
0.54 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
177 |
0.63 |
Binding ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
53 |
0.68 |
Binding ≤ 10μM
|
|
GRIK3-2-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
10 |
0.75 |
Binding ≤ 10μM
|
|
GRIK4-2-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
7 |
0.76 |
Binding ≤ 10μM
|
|
GRIK5-2-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
7 |
0.76 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
7510 |
0.48 |
Functional ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
700 |
0.57 |
Functional ≤ 10μM
|
|
Z50597-4-O |
Rattus Norvegicus (cluster #4 Of 5), Other |
Other |
18 |
0.72 |
Binding ≤ 10μM
|
|
Z50466-5-O |
Trypanosoma Cruzi (cluster #5 Of 8), Other |
Other |
3000 |
0.52 |
Functional ≤ 10μM
|
|
Z50597-6-O |
Rattus Norvegicus (cluster #6 Of 12), Other |
Other |
3700 |
0.51 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-0.81 |
5.21 |
-88.22 |
2 |
5 |
-1 |
97 |
212.225 |
4 |
↓
|
|
|
|
Analogs
-
12504088
-
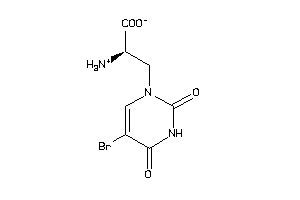
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
35 |
0.70 |
Binding ≤ 10μM
|
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
92 |
0.66 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
61 |
0.67 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
101 |
0.65 |
Binding ≤ 10μM
|
|
GRIA3-4-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
434 |
0.59 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
457 |
0.59 |
Binding ≤ 10μM
|
|
GRIA4-4-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
656 |
0.58 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
15 |
0.73 |
Binding ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
9 |
0.75 |
Binding ≤ 10μM
|
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
2800 |
0.52 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.22 |
-4.87 |
-39.23 |
4 |
7 |
0 |
122 |
278.062 |
3 |
↓
|
|
|
|
Analogs
-
13473523
-
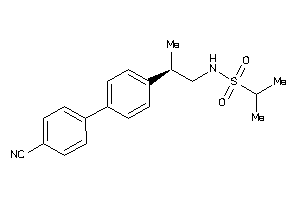
Draw
Identity
99%
90%
80%
70%
Vendors
And 1 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-4-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
5650 |
0.31 |
Binding ≤ 10μM
|
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1440 |
0.34 |
Binding ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1660 |
0.34 |
Binding ≤ 10μM
|
|
GRIA4-5-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
290 |
0.38 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
559 |
0.36 |
Functional ≤ 10μM
|
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
420 |
0.37 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.36 |
7.99 |
-14.33 |
1 |
4 |
0 |
70 |
342.464 |
6 |
↓
|
|
|
|
Analogs
-
8815
-
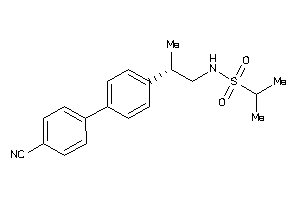
Draw
Identity
99%
90%
80%
70%
Vendors
And 2 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-4-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
5650 |
0.31 |
Binding ≤ 10μM
|
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1440 |
0.34 |
Binding ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
1660 |
0.34 |
Binding ≤ 10μM
|
|
GRIA4-5-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
290 |
0.38 |
Binding ≤ 10μM
|
|
GRIA2-4-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #4 Of 4), Eukaryotic |
Eukaryotes |
559 |
0.36 |
Functional ≤ 10μM
|
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
420 |
0.37 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.36 |
7.98 |
-14.96 |
1 |
4 |
0 |
70 |
342.464 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 145 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
AA3R-6-E |
Adenosine Receptor A3 (cluster #6 Of 6), Eukaryotic |
Eukaryotes |
613 |
0.87 |
Binding ≤ 10μM
|
|
EAA3-2-E |
Excitatory Amino Acid Transporter 3 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
51 |
1.02 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
1360 |
0.82 |
Binding ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
940 |
0.84 |
Binding ≤ 10μM
|
|
GRIA3-3-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
500 |
0.88 |
Binding ≤ 10μM
|
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
868 |
0.85 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
701 |
0.86 |
Binding ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
1106 |
0.83 |
Binding ≤ 10μM
|
|
GRIK3-2-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
789 |
0.85 |
Binding ≤ 10μM
|
|
GRIK4-2-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
700 |
0.86 |
Binding ≤ 10μM
|
|
GRIK5-2-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
700 |
0.86 |
Binding ≤ 10μM
|
|
GRM1-3-E |
Metabotropic Glutamate Receptor 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
570 |
0.87 |
Binding ≤ 10μM
|
|
GRM2-1-E |
Metabotropic Glutamate Receptor 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7400 |
0.72 |
Binding ≤ 10μM
|
|
GRM3-1-E |
Metabotropic Glutamate Receptor 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
3000 |
0.77 |
Binding ≤ 10μM
|
|
GRM4-3-E |
Metabotropic Glutamate Receptor 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
2400 |
0.79 |
Binding ≤ 10μM
|
|
GRM5-4-E |
Metabotropic Glutamate Receptor 5 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
4100 |
0.75 |
Binding ≤ 10μM
|
|
GRM6-1-E |
Metabotropic Glutamate Receptor 6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
7600 |
0.72 |
Binding ≤ 10μM
|
|
GRM8-1-E |
Metabotropic Glutamate Receptor 8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
3400 |
0.77 |
Binding ≤ 10μM
|
|
NMDZ1-4-E |
Glutamate (NMDA) Receptor Subunit Zeta 1 (cluster #4 Of 6), Eukaryotic |
Eukaryotes |
450 |
0.89 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
6160 |
0.73 |
Functional ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
6200 |
0.73 |
Functional ≤ 10μM
|
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
3660 |
0.76 |
Functional ≤ 10μM
|
|
GRM1-4-E |
Metabotropic Glutamate Receptor 1 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
10000 |
0.70 |
Functional ≤ 10μM
|
|
GRM2-1-E |
Metabotropic Glutamate Receptor 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
8500 |
0.71 |
Functional ≤ 10μM
|
|
GRM3-1-E |
Metabotropic Glutamate Receptor 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
9000 |
0.71 |
Functional ≤ 10μM
|
|
GRM4-1-E |
Metabotropic Glutamate Receptor 4 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
6100 |
0.73 |
Functional ≤ 10μM
|
|
GRM5-5-E |
Metabotropic Glutamate Receptor 5 (cluster #5 Of 5), Eukaryotic |
Eukaryotes |
4900 |
0.74 |
Functional ≤ 10μM
|
|
GRM6-1-E |
Metabotropic Glutamate Receptor 6 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
7600 |
0.72 |
Functional ≤ 10μM
|
|
GRM8-1-E |
Metabotropic Glutamate Receptor 8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
9500 |
0.70 |
Functional ≤ 10μM
|
|
NMDE1-1-E |
Glutamate [NMDA] Receptor Subunit Epsilon 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1800 |
0.80 |
Functional ≤ 10μM
|
|
NMDZ1-1-E |
Glutamate (NMDA) Receptor Subunit Zeta 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
1800 |
0.80 |
Functional ≤ 10μM
|
|
Z104302-3-O |
Glutamate NMDA Receptor (cluster #3 Of 7), Other |
Other |
870 |
0.85 |
Binding ≤ 10μM
|
|
Z50597-3-O |
Rattus Norvegicus (cluster #3 Of 5), Other |
Other |
170 |
0.95 |
Binding ≤ 10μM
|
|
Z50597-7-O |
Rattus Norvegicus (cluster #7 Of 12), Other |
Other |
1300 |
0.82 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.25 |
1.74 |
-73.91 |
3 |
5 |
-1 |
108 |
146.122 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 120 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
MURI-2-B |
Glutamate Racemase (cluster #2 Of 2), Bacterial |
Bacteria |
5800 |
0.73 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
1362 |
0.82 |
Binding ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
940 |
0.84 |
Binding ≤ 10μM
|
|
GRIA3-3-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
500 |
0.88 |
Binding ≤ 10μM
|
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 5), Eukaryotic |
Eukaryotes |
868 |
0.85 |
Binding ≤ 10μM
|
|
GRIK1-3-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
701 |
0.86 |
Binding ≤ 10μM
|
|
GRIK2-2-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
1106 |
0.83 |
Binding ≤ 10μM
|
|
GRIK3-2-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
789 |
0.85 |
Binding ≤ 10μM
|
|
GRIK4-2-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
400 |
0.90 |
Binding ≤ 10μM
|
|
GRIK5-2-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
400 |
0.90 |
Binding ≤ 10μM
|
|
GRM1-3-E |
Metabotropic Glutamate Receptor 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
4900 |
0.74 |
Binding ≤ 10μM
|
|
GRM2-1-E |
Metabotropic Glutamate Receptor 2 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
290 |
0.92 |
Binding ≤ 10μM
|
|
GRM3-1-E |
Metabotropic Glutamate Receptor 3 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1900 |
0.80 |
Binding ≤ 10μM
|
|
GRM4-3-E |
Metabotropic Glutamate Receptor 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
9800 |
0.70 |
Binding ≤ 10μM
|
|
GRM5-4-E |
Metabotropic Glutamate Receptor 5 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
3100 |
0.77 |
Binding ≤ 10μM
|
|
GRM6-1-E |
Metabotropic Glutamate Receptor 6 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
4900 |
0.74 |
Binding ≤ 10μM
|
|
NMDE1-4-E |
Glutamate [NMDA] Receptor Subunit Epsilon 1 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
200 |
0.94 |
Binding ≤ 10μM
|
|
NMDE2-4-E |
Glutamate [NMDA] Receptor Subunit Epsilon 2 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
200 |
0.94 |
Binding ≤ 10μM
|
|
NMDE3-3-E |
Glutamate [NMDA] Receptor Subunit Epsilon 3 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
200 |
0.94 |
Binding ≤ 10μM
|
|
NMDE4-2-E |
Glutamate [NMDA] Receptor Subunit Epsilon 4 (cluster #2 Of 6), Eukaryotic |
Eukaryotes |
200 |
0.94 |
Binding ≤ 10μM
|
|
GRIA1-3-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
214 |
0.93 |
Functional ≤ 10μM
|
|
GRIA2-3-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #3 Of 4), Eukaryotic |
Eukaryotes |
214 |
0.93 |
Functional ≤ 10μM
|
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 2), Eukaryotic |
Eukaryotes |
214 |
0.93 |
Functional ≤ 10μM
|
|
GRIA4-2-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
214 |
0.93 |
Functional ≤ 10μM
|
|
GRM1-4-E |
Metabotropic Glutamate Receptor 1 (cluster #4 Of 5), Eukaryotic |
Eukaryotes |
1050 |
0.84 |
Functional ≤ 10μM
|
|
GRM8-1-E |
Metabotropic Glutamate Receptor 8 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
8000 |
0.71 |
Functional ≤ 10μM
|
|
Z104302-3-O |
Glutamate NMDA Receptor (cluster #3 Of 7), Other |
Other |
80 |
0.99 |
Binding ≤ 10μM |
|
Z50597-7-O |
Rattus Norvegicus (cluster #7 Of 12), Other |
Other |
2500 |
0.78 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
-3.25 |
1.72 |
-74 |
3 |
5 |
-1 |
108 |
146.122 |
4 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
And 15 More
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA1-2-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
340 |
0.53 |
Binding ≤ 10μM |
|
GRIA2-2-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
340 |
0.53 |
Binding ≤ 10μM |
|
GRIA3-2-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #2 Of 4), Eukaryotic |
Eukaryotes |
340 |
0.53 |
Binding ≤ 10μM |
|
GRIA4-2-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #2 Of 5), Eukaryotic |
Eukaryotes |
340 |
0.53 |
Binding ≤ 10μM |
|
GRIK1-2-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #2 Of 3), Eukaryotic |
Eukaryotes |
2400 |
0.46 |
Binding ≤ 10μM |
|
GRIK2-1-E |
Glutamate Receptor Ionotropic Kainate 2 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2400 |
0.46 |
Binding ≤ 10μM |
|
GRIK3-1-E |
Glutamate Receptor Ionotropic Kainate 3 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
2400 |
0.46 |
Binding ≤ 10μM |
|
GRIK4-1-E |
Glutamate Receptor Ionotropic Kainate 4 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2400 |
0.46 |
Binding ≤ 10μM |
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
2400 |
0.46 |
Binding ≤ 10μM |
|
NMDZ1-1-E |
Glutamate (NMDA) Receptor Subunit Zeta 1 (cluster #1 Of 6), Eukaryotic |
Eukaryotes |
7900 |
0.42 |
Binding ≤ 10μM |
|
GRIA1-1-E |
Glutamate Receptor Ionotropic, AMPA 1 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
600 |
0.51 |
Functional ≤ 10μM
|
|
GRIA2-1-E |
Glutamate Receptor Ionotropic, AMPA 2 (cluster #1 Of 4), Eukaryotic |
Eukaryotes |
600 |
0.51 |
Functional ≤ 10μM
|
|
GRIA3-1-E |
Glutamate Receptor Ionotropic, AMPA 3 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
600 |
0.51 |
Functional ≤ 10μM
|
|
GRIA4-1-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #1 Of 3), Eukaryotic |
Eukaryotes |
600 |
0.51 |
Functional ≤ 10μM
|
|
GRIK1-1-E |
Glutamate Receptor Ionotropic Kainate 1 (cluster #1 Of 2), Eukaryotic |
Eukaryotes |
1660 |
0.48 |
Functional ≤ 10μM
|
|
GRIK5-1-E |
Glutamate Receptor Ionotropic Kainate 5 (cluster #1 Of 1), Eukaryotic |
Eukaryotes |
1660 |
0.48 |
Functional ≤ 10μM
|
|
Z104302-1-O |
Glutamate NMDA Receptor (cluster #1 Of 7), Other |
Other |
5600 |
0.43 |
Binding ≤ 10μM
|
|
Z50425-3-O |
Plasmodium Falciparum (cluster #3 Of 22), Other |
Other |
10000 |
0.41 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
0.26 |
0.17 |
-38.85 |
1 |
8 |
-1 |
138 |
231.147 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.22 |
1.62 |
-52.45 |
1 |
8 |
1 |
131 |
231.147 |
1 |
↓
|
|
Hi
High (pH 8-9.5)
|
0.22 |
1.62 |
-53.61 |
1 |
8 |
1 |
131 |
231.147 |
1 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
551 |
0.49 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.17 |
5.34 |
-8.87 |
1 |
3 |
0 |
46 |
267.394 |
4 |
↓
|
|
|
|
Analogs
-
13835063
-
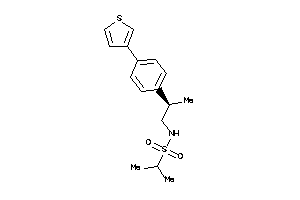
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
660 |
0.41 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.20 |
6.65 |
-12.23 |
1 |
3 |
0 |
46 |
323.483 |
6 |
↓
|
|
|
|
Analogs
-
26120
-
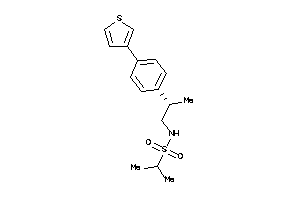
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
660 |
0.41 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
4.20 |
6.65 |
-11.21 |
1 |
3 |
0 |
46 |
323.483 |
6 |
↓
|
|
|
|
Analogs
Draw
Identity
99%
90%
80%
70%
Vendors
Clustered Target Annotations
| Code |
Organism Class |
Affinity (nM) |
LE (kcal/mol/atom) |
Type |
|
GRIA4-3-E |
Glutamate Receptor Ionotropic, AMPA 4 (cluster #3 Of 3), Eukaryotic |
Eukaryotes |
1740 |
0.45 |
Functional ≤ 10μM
|
Physical Representations
|
Type
pH range
|
xlogP
|
Des A‑Pol
Apolar desolvation
(kcal/mol)
|
Des Pol
Polar desolvation
(kcal/mol)
|
H Don
H-bond donors
|
H Acc
H-bond acceptors
|
Chg
Net charge
|
tPSA
(Ų)
|
MWT
Molecular weight
(g/mol)
|
RB
Rotatable bonds
|
DL |
|
Ref
Reference (pH 7)
|
3.17 |
5.83 |
-10.86 |
1 |
3 |
0 |
46 |
267.394 |
4 |
↓
|
|